MA_MAT3VR3351
PREDICTED INTERACTION BETWEEN XRCC5 (1-587) AND HIF1A (789-826)
There are no experimental data to verify the accuracy of this computed structure model. See Model Confidence metrics below for all regions of the polypeptide chain.
- ModelArchive: ma-t3vr3-351
- Released in ModelArchive: 2022-12-21
- Organism(s): Homo sapiens
- UniProtKB: P13010 Q16665
Model Confidence
- pLDDT (global): 77.629
- pLDDT (local):
Model Confidence
- Very high (pLDDT > 90)
- Confident (70 < pLDDT ≤ 90)
- Low (50 < pLDDT ≤ 70)
- Very low (pLDDT ≤ 50)
Computed Structure Models provide per-residue confidence score (pLDDT) between 0 and 100. Some regions below 50 pLDDT may be unstructured in isolation.
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Homo sapiens (Human) XRCC5 (P13010; 1 - 587) | 587 | Homo sapiens | Mutation(s): 0 EC: 5.6.2.4 (UniProt), 4.2.99.18 (UniProt) | 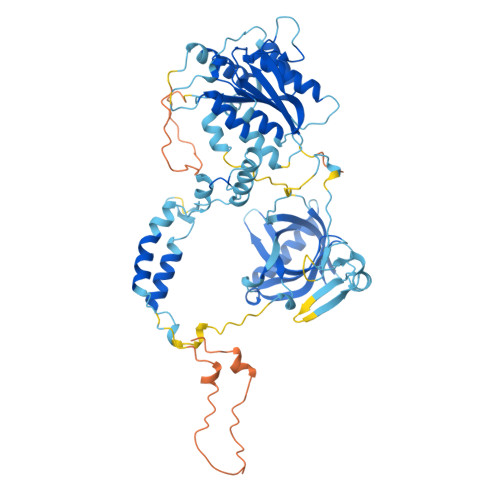 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P13010 (Homo sapiens) Go to UniProtKB: P13010 | |||||
PHAROS: P13010 GTEx: ENSG00000079246 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Homo sapiens (Human) HIF1A (Q16665; 789 - 826) | 38 | Homo sapiens | Mutation(s): 0 | 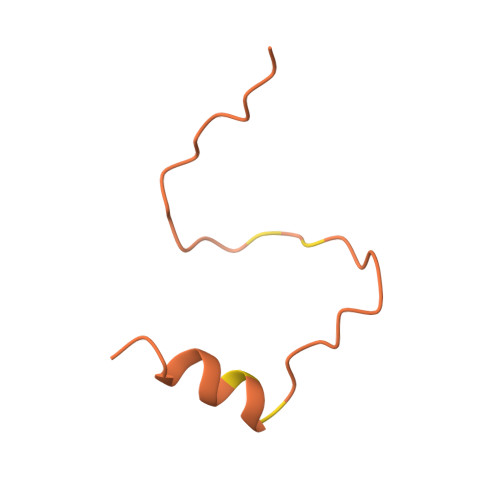 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for Q16665 (Homo sapiens) Go to UniProtKB: Q16665 | |||||
PHAROS: Q16665 GTEx: ENSG00000100644 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
Sequence AnnotationsExpand | |||||
| |||||














