AF_AFQ46824F1
COMPUTED STRUCTURE MODEL OF INNER MEMBRANE PROTEIN YGFX
There are no experimental data to verify the accuracy of this computed structure model. See Model Confidence metrics below for all regions of the polypeptide chain.
- AlphaFold DB: AF-Q46824-F1
- Released in AlphaFold DB: 2021-07-01
Last Modified in AlphaFold DB: 2022-09-30 - Organism(s): Escherichia coli K-12
- UniProtKB: Q46824
Model Confidence
- pLDDT (global): 78.27
- pLDDT (local):
Model Confidence
- Very high (pLDDT > 90)
- Confident (70 < pLDDT ≤ 90)
- Low (50 < pLDDT ≤ 70)
- Very low (pLDDT ≤ 50)
Computed Structure Models provide per-residue confidence score (pLDDT) between 0 and 100. Some regions below 50 pLDDT may be unstructured in isolation.
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Inner membrane protein YgfX | 135 | Escherichia coli K-12 | Mutation(s): 0 Gene Names: ygfX | 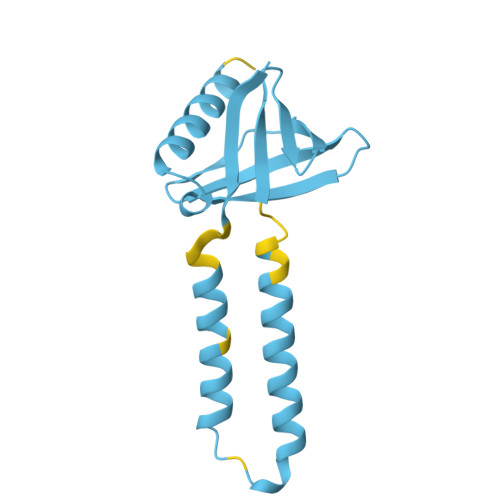 | |
UniProt | |||||
Find proteins for Q46824 (Escherichia coli (strain K12)) Explore Q46824 Go to UniProtKB: Q46824 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q46824 | ||||
Sequence AnnotationsExpand | |||||
| |||||














