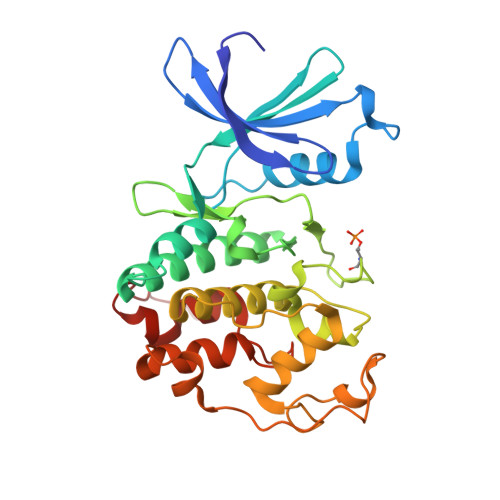
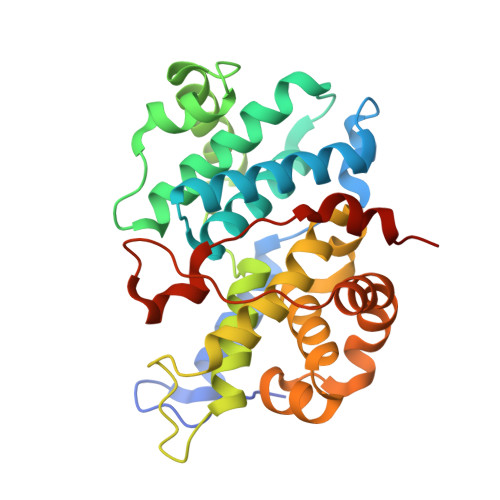
Nanoscale Direct-to-Biology Optimization of Cdk2 Inhibitors.
Douthwaite, J.L., Houde, D.J., Pardo, E., Moran, M., Baird, J., Meyer, S.R., Mahjour, B., Zhao, Q., Larrow, J.F., Juang, Y.P., Holliday, M.J., Han, C., Kelley, B., Dunstan, D., Billings, K., Mader, M.M., Taylor, A.M., Sexton, J.Z., Boezio, A.A., Cernak, T.(2026) J Med Chem 69: 9142-9162
- PubMed: 41954971 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.5c03614
- Primary Citation Related Structures:
9OB2, 9OB3, 9OB4, 9OB5, 9OB6 - PubMed Abstract:
Modern hit-to-lead optimization winnows down vast chemical spaces of virtual compounds into a selection of potent and selective compounds that can be further profiled with in vitro assays. Today, miniaturized chemical synthesis can be performed in high-throughput, shifting the bottleneck to compound purification. Direct-to-biology (D2B) approaches seek to overcome this hurdle by omitting the purification step and submitting reaction mixtures directly to bioassays. Here, we explore nanoscale hit-to-lead optimization through the multistep synthesis of a library of Cdk2/CycE inhibitors, utilizing ultrahigh-throughput experimentation (ultraHTE) in 1,536-well plates. Library performance is assessed by D2B in functional biochemical, bioaffinity, and X-ray crystallographic assays. A selection of potent lead Cdk2/CycE inhibitors identified by D2B was submitted to a phenotypic cell painting assay, which showed cell cycle arrest at G 0 , consistent with Cdk2 inhibition. This miniaturized workflow allows the upper tiers of a typical optimization screening cascade to be performed in a single experiment.
- Department of Medicinal Chemistry, College of Pharmacy, University of Michigan, Ann Arbor, Michigan 48109, United States.
Organizational Affiliation: