Chemically-induced degradation of the endoplasmic-reticulum stress sensor IRE1 by a VHL-recruiting chimera.
Du, J., Villemure, E., Johnson, M., Azumaya, C., Gaspar, C.J., Marsters, S., Lawrence, D., Foster, S., Rohou, A., Cheung, T.K., Rose, C.M., Garner, T., Ro, S., Clark, K., Beresini, M.H., Braun, M.G., Rudolph, J., Hsu, P., Ashkenazi, A.(2025) Nat Commun 16: 11445-11445
- PubMed: 41381506 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-66382-8
- Primary Citation Related Structures:
9N88 - PubMed Abstract:
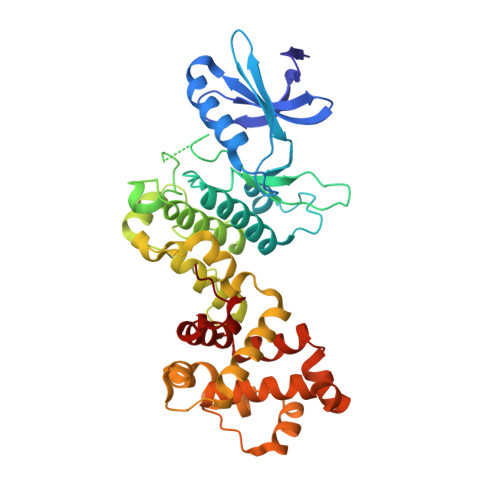
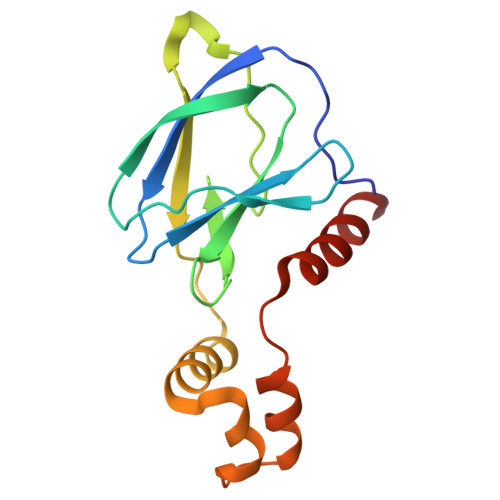
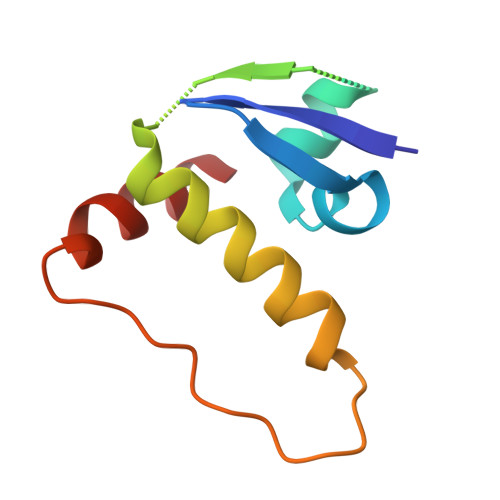
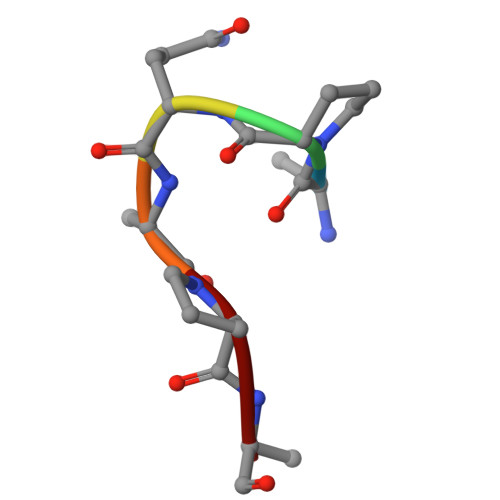
The endoplasmic-reticulum (ER) transmembrane protein IRE1 mitigates ER stress through kinase-endoribonuclease and scaffolding activities. Cancer cells often co-opt IRE1 to facilitate growth. An IRE1-RNase inhibitor has entered clinical trials; however, recent work uncovered a significant nonenzymatic IRE1 dependency in cancer. To fully disrupt IRE1, we describe a proteolysis-targeting chimera (G6374) that couples an IRE1-kinase ligand to a compound that binds the ubiquitin Cullin-RING Ligase (CRL) substrate receptor, VHL. G6374 induces a stable, cooperative interaction between IRE1 and VHL, driving K48-linked ubiquitination on two principal lysine residues in the IRE1-kinase domain and inducing proteasomal IRE1 degradation. Cryogenic electron microscopy and mutagenesis studies reveal a 2:2 IRE1:VHL ternary-complex topology and critical interactional features, informing future designs. G6374 blocks growth of IRE1-dependent cancer cells irrespective of their dependency mode, while sparing IRE1-independent cells. We provide a proof-of-concept for VHL-based degradation of an ER-transmembrane protein, advancing strategies to fully disrupt IRE1.
- Department of Research Oncology, Genentech, Inc., South San Francisco, CA, USA.
Organizational Affiliation: