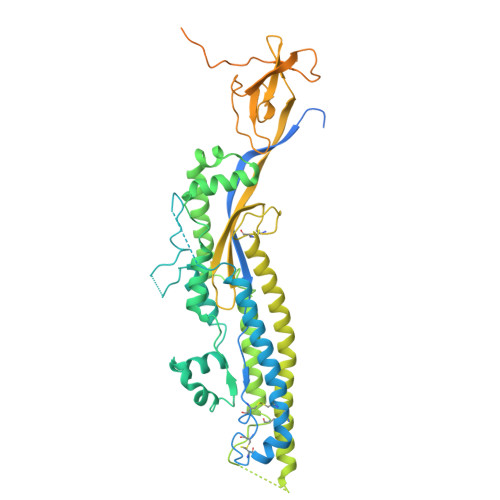
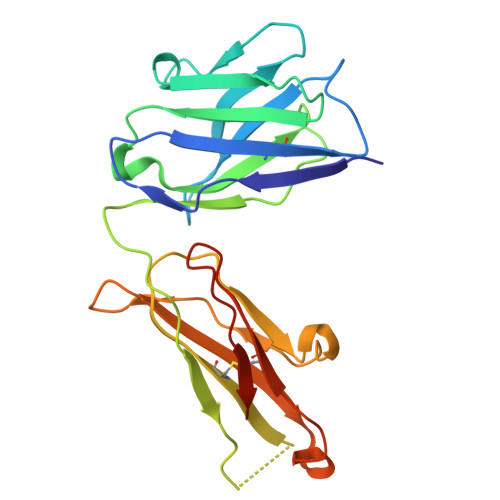
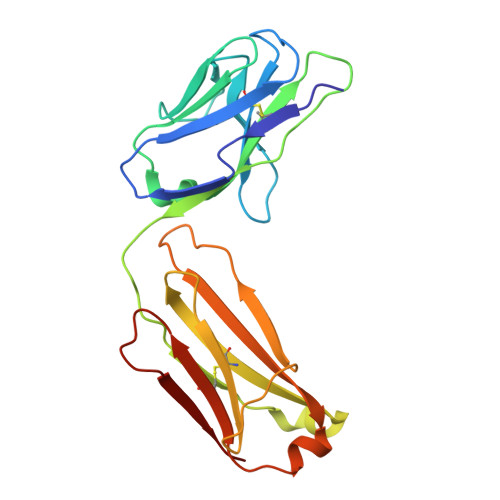
Clonotype-enriched somatic hypermutations drive affinity maturation of a public human antibody targeting an occluded sarbecovirus epitope.
Rao, V.N., Sapse, I.A., Cohn, H., Yoo, D.K., Tong, P., Clark, J.J., Bozarth, B., Chen, Y., Srivastava, K., Singh, G., Krammer, F., Simon, V., Wesemann, D.R., Bajic, G., Coelho, C.H.(2025) Cell Rep 44: 116122-116122
- PubMed: 40803328 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.celrep.2025.116122
- Primary Citation Related Structures:
9MPW - PubMed Abstract:
Investigating public antibodies that recognize conserved epitopes is critical for vaccine development. Identifying somatic hypermutations (SHMs) that enhance antigen affinity in these public antibodies is key to guiding vaccine design for better protection against pathogens. We propose that affinity-enhancing SHMs are selectively enriched in public antibody clonotypes, surpassing the background frequency seen in antibodies carrying the same V genes but with different epitope specificities. Using M15, a human IGHV4-59/IGKV3-20 public antibody as a model, we compare SHM signatures in antibodies that use the same V genes but recognize other epitopes. We identified clonotype-enriched mutations in the light chain of M15 and showed that, in combination, these SHMs enhance binding to a previously uncharacterized Sarbecovirus epitope, with antibody responses to it increasing after sequential vaccination. Our findings identify convergence and clonotype enrichment as features of affinity-enhancing SHMs in public antibodies, which can help guide vaccine design aimed at eliciting such antibodies.
- Department of Microbiology, Icahn School of Medicine at Mount Sinai, New York, NY, USA; Center for Vaccine Research and Pandemic Preparedness, Icahn School of Medicine at Mount Sinai, New York, NY, USA; Graduate School of Biomedical Sciences, Icahn School of Medicine at Mount Sinai, New York, NY, USA.
Organizational Affiliation: