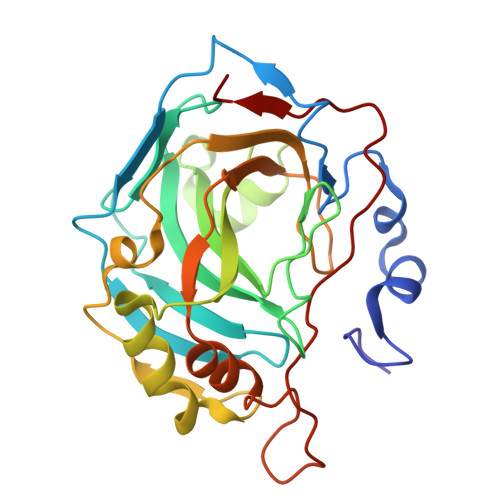
Exploring the Polypharmacological Potential of PCI-27483: A Selective Inhibitor of Carbonic Anhydrases IX and XII.
D'Agostino, I., Bonardi, A., Ferraroni, M., Gratteri, P., Angeli, A., Supuran, C.T.(2024) ACS Med Chem Lett 15: 2042-2045
- PubMed: 39563799 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsmedchemlett.4c00443
- Primary Citation Related Structures:
9GOO - PubMed Abstract:
PCI-27483, originally developed as a potent and selective inhibitor of the serine protease Factor VIIa (FVIIa) in complex with tissue factor (TF), has demonstrated significant promise in cancer therapy. In addition to its primary mechanism of action, the presence of a sulfonamide moiety in the PCI-27483 structure suggests further activities through the inhibition of carbonic anhydrases (CAs), particularly the tumor-associated human (h)CA isoforms hCA IX and XII. This study investigates the inhibitory activity of PCI-27483 against the complete panel of active hCAs, highlighting its polypharmacological potential in cancer treatment. X-ray crystallography and molecular docking studies elucidated the structural features underlying its selective inhibitory activity toward hCA IX and XII, offering insights into its dual-targeting pathway.
- Department of Pharmacy, University of Pisa, Via Bonanno 6, 56126 Pisa, Italy.
Organizational Affiliation: