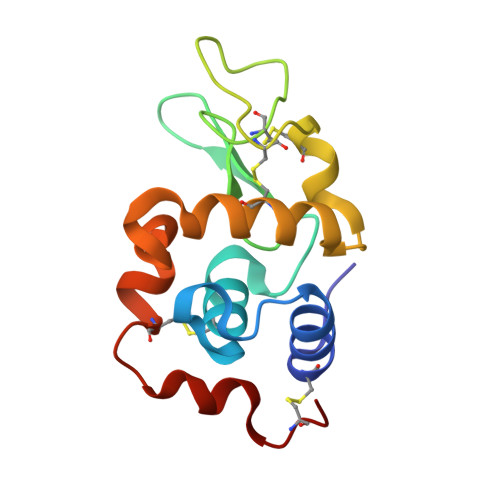
Body temperature protein X-ray crystallography at 37 °C: a rhenium protein complex seeking a physiological condition structure.
Jacobs, F.J.F., Helliwell, J.R., Brink, A.(2024) Chem Commun (Camb) 60: 14030-14033
- PubMed: 39382205 Search on PubMed
- DOI: https://doi.org/10.1039/d4cc04245j
- Primary Citation Related Structures:
9GHX - PubMed Abstract:
The retention of the covalent binding of an organometalllic rhenium complex as a model for a technetium-99m imaging agent, to a protein at physiological body temperature 37 °C is described. Detailed structure comparisons are made to the related 100 K crystal structure. The generality of the need for this sort of analytical procedure for guiding ligand lead compound discovery is emphasised.
- Department of Chemistry, University of the Free State, Nelson Mandela Drive, Bloemfontein 9301, South Africa.
Organizational Affiliation: