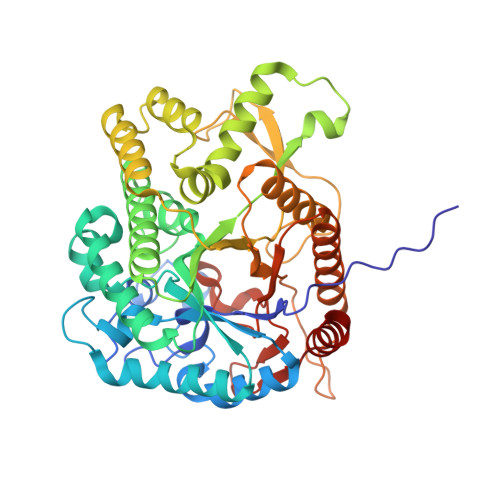
Structural studies of beta-glucosidase from the thermophilic bacterium Caldicellulosiruptor saccharolyticus.
Sotiropoulou, A.I., Hatzinikolaou, D.G., Chrysina, E.D.(2024) Acta Crystallogr D Struct Biol 80: 733-743
- PubMed: 39361356 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2059798324009252
- Primary Citation Related Structures:
9GCI, 9GCJ - PubMed Abstract:
β-Glucosidase from the thermophilic bacterium Caldicellulosiruptor saccharolyticus (Bgl1) has been denoted as having an attractive catalytic profile for various industrial applications. Bgl1 catalyses the final step of in the decomposition of cellulose, an unbranched glucose polymer that has attracted the attention of researchers in recent years as it is the most abundant renewable source of reduced carbon in the biosphere. With the aim of enhancing the thermostability of Bgl1 for a broad spectrum of biotechnological processes, it has been subjected to structural studies. Crystal structures of Bgl1 and its complex with glucose were determined at 1.47 and 1.95 Å resolution, respectively. Bgl1 is a member of glycosyl hydrolase family 1 (GH1 superfamily, EC 3.2.1.21) and the results showed that the 3D structure of Bgl1 follows the overall architecture of the GH1 family, with a classical (β/α) 8 TIM-barrel fold. Comparisons of Bgl1 with sequence or structural homologues of β-glucosidase reveal quite similar structures but also unique structural features in Bgl1 with plausible functional roles.
- Institute of Chemical Biology, National Hellenic Research Foundation, 48 Vassileos Constantinou Avenue, 116 35 Athens, Greece.
Organizational Affiliation: