Single Acetylation-mimetic Mutation in TDP-43 Nuclear Localization Signal Disrupts Importin alpha 1/ beta Signaling.
Ko, Y.H., Lokareddy, R.K., Doll, S.G., Yeggoni, D.P., Girdhar, A., Mawn, I., Klim, J.R., Rizvi, N.F., Meyers, R., Gillilan, R.E., Guo, L., Cingolani, G.(2024) J Mol Biology 436: 168751-168751
- PubMed: 39181183 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2024.168751
- Primary Citation Related Structures:
9B4Y - PubMed Abstract:
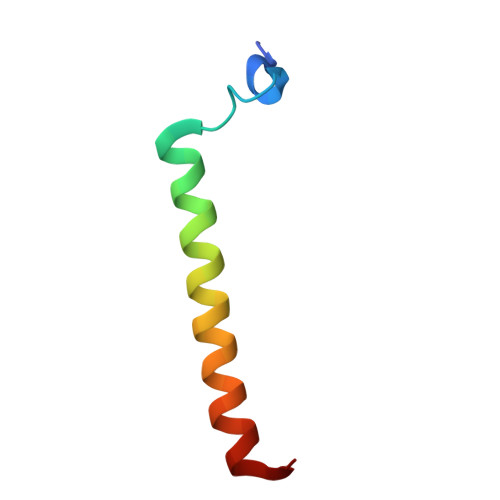
Cytoplasmic aggregation of the TAR-DNA binding protein of 43 kDa (TDP-43) is the hallmark of sporadic amyotrophic lateral sclerosis (ALS). Most ALS patients with TDP-43 aggregates in neurons and glia do not have mutations in the TDP-43 gene but contain aberrantly post-translationally modified TDP-43. Here, we found that a single acetylation-mimetic mutation (K82Q) near the TDP-43 minor Nuclear Localization Signal (NLS) box, which mimics a post-translational modification identified in an ALS patient, can lead to TDP-43 mislocalization to the cytoplasm and irreversible aggregation. We demonstrate that the acetylation mimetic disrupts binding to importins, halting nuclear import and preventing importin α1/β anti-aggregation activity. We propose that perturbations near the NLS are an additional mechanism by which a cellular insult other than a genetically inherited mutation leads to TDP-43 aggregation and loss of function. Our findings are relevant to deciphering the molecular etiology of sporadic ALS.
- Dept. of Biochemistry and Molecular Genetics, The University of Alabama at Birmingham, 1825 University Blvd, Birmingham, AL 35294, USA.
Organizational Affiliation: