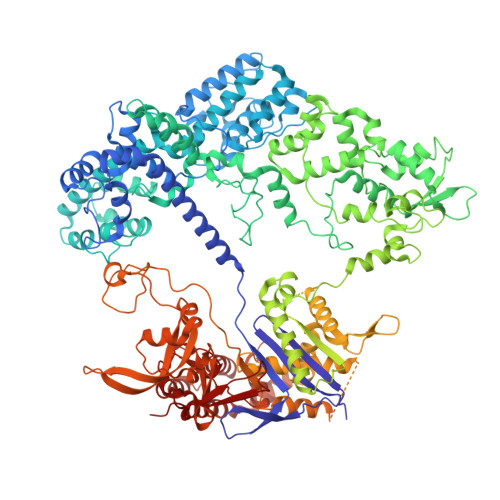
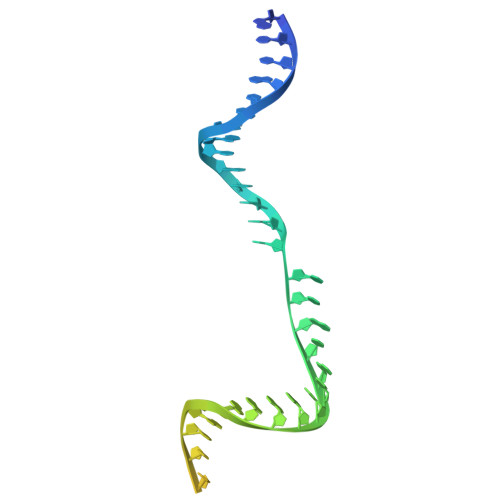
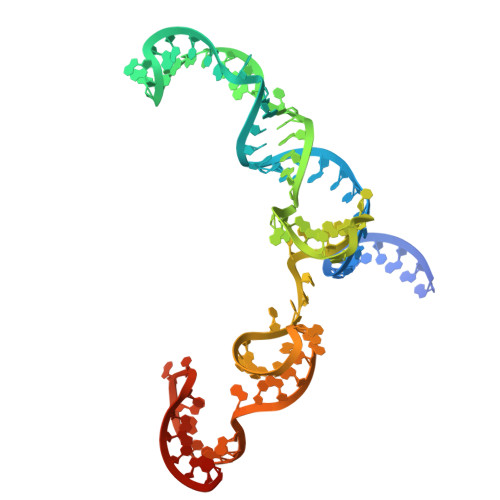
Improving the efficiency of high-fidelity Cas9 by enhancing PAM-distal interactions.
Zheng, R., Lu, Z., Wei, R., Shin, Y.C., Du, J., Zhang, Q., Li, J., Wang, X., Wei, Y., Liu, B., Chen, Y., Ding, L., Zhang, H., Chen, H., Huang, J., Ma, L.(2026) Nat Struct Mol Biol 33: 590-602
- PubMed: 41851507 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-026-01753-3
- Primary Citation Related Structures:
9K4C, 9K4D, 9K4E, 9K4F, 9K4H, 9W7Q, 9W7T, 9W7U, 9W7V, 9W9D, 9WA9, 9WAA, 9WAW - PubMed Abstract:
Engineering CRISPR enzymes for high fidelity often impairs cleavage activity. Meanwhile, a mechanistic understanding of why high-fidelity mutations reduce Cas9's cleavage activity remains unclear, presenting a challenge in balancing nuclease specificity and efficiency for clinical applications. In this study, we show that extending the spacer region to 21 or 22 nucleotides restores the impaired cleavage activity of SuperFi-Cas9, a high-fidelity Cas9 variant with 7 mutations in the RuvC domain at the protospacer adjacent motif (PAM)-distal region. Cryo-electron microscopy structures and mutational analyses reveal that the negatively charged mutations in a protruding loop of the RuvC domain create repulsive forces that destabilize the nuclease-single guide (sg)RNA-DNA complex. Spacer extension enhances interactions in the PAM-distal region, effectively restoring cleavage activity and balancing editing efficiency with specificity. In addition, we develop a deep learning model, AIdit-SuperFi, to predict optimal sgRNA length for high-fidelity genome editing. Our findings introduce a straightforward strategy to enhance CRISPR complex stability and provide mechanistic insights into the impaired cleavage activity of engineered high-fidelity Cas9, presenting a pathway toward precise and efficient genome editing and clinical translation of CRISPR technologies.
- Westlake Laboratory, Hangzhou, China.
Organizational Affiliation: