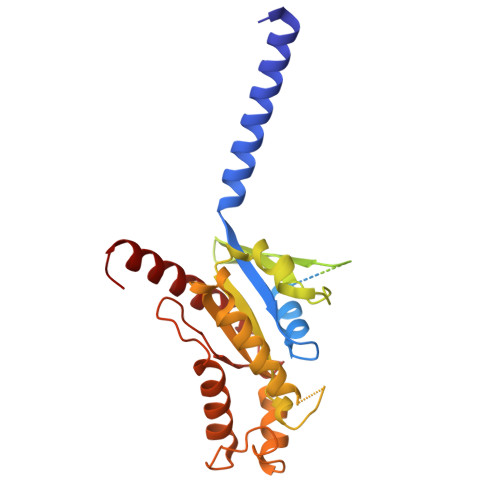
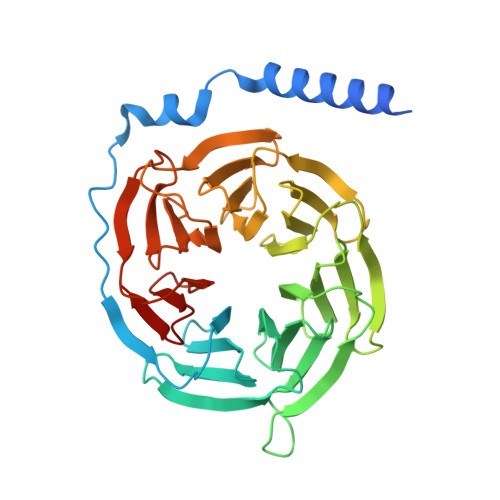
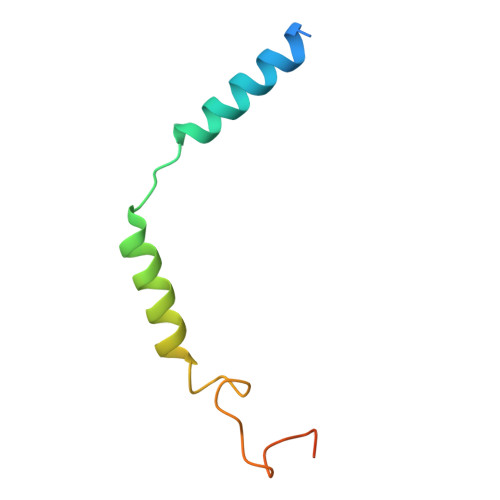
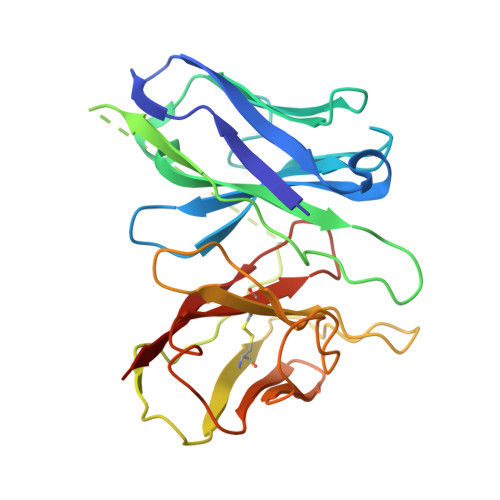
Pathway-selective 5-HT 1A R agonist as a rapid antidepressant strategy.
Wang, C., Zhang, N., Shao, Y., Li, T., Zhang, M., Gao, M., Liang, Y., Wang, Y., Xue, T., Shi, Y., Chen, H., Cao, C.(2025) Cell 188: 7222-7237.e24
- PubMed: 41232528 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2025.10.022
- Primary Citation Related Structures:
9VJ5, 9VJ6, 9VJE, 9VJF, 9VJG, 9VMY, 9VNF - PubMed Abstract:
Presynaptic 5-HT 1A R autoreceptors predominantly signal through G i3 protein, mediating feedback inhibition that hampers the therapeutic efficacy of conventional antidepressants. By contrast, postsynaptic heteroreceptors mainly couple to G o , which promotes antidepressant responses. However, selectively activating heteroreceptors while bypassing the negative feedback induced by autoreceptors remains a significant challenge. Here, we characterized the G i/o subtype signaling profiles of 5-HT 1A R and determined its structures in complex with six agonists and three distinct G i/o family proteins: G oA , G i3 , and G z . Combined with functional analysis, we elucidated the mechanisms underlying diverse agonist recognition modes and G i/o subtype signaling selectivity of 5-HT 1A R. Furthermore, we designed a pathway-selective agonist, TMU4142, which exhibits high G oA activity while minimizing G i3 activation. Remarkably, TMU4142 demonstrated rapid antidepressant-like effects in a mouse model of depression. Collectively, these findings suggest that distinguishing heteroreceptors from autoreceptors based on their distinct downstream G i/o signaling pathways could be a promising strategy to develop fast-acting antidepressants.
- Institute of Health and Medicine, Hefei Comprehensive National Science Center; State Key Laboratory of Immune Response and Immunotherapy, Division of Life Sciences and Medicine, University of Science and Technology of China, Hefei, China. Electronic address: cywang@ihm.ac.cn.
Organizational Affiliation: