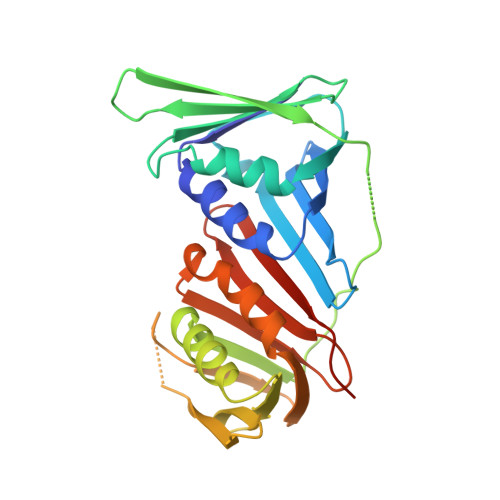
Identification of a PCNA-binding motif in human translesion DNA polymerase REV1 and structural basis of its interaction with PCNA.
Hishiki, A., Hoshino, N., Okawara, K., Fuchigami, S., Hara, K., Hashimoto, H.(2025) J Biochem 178: 315-324
- PubMed: 40888629 Search on PubMed
- DOI: https://doi.org/10.1093/jb/mvaf054
- Primary Citation Related Structures:
9VGW - PubMed Abstract:
REV1 is a eukaryotic error-prone DNA polymerase belonging to the Y-family, with a central role in translesion DNA synthesis (TLS) to continue DNA replication even in the presence of DNA damage in the template strand. TLS is stimulated by mono-ubiquitination of proliferating cell nuclear antigen (PCNA), a toroidal-shaped protein functioning as a scaffold for DNA polymerases and repair enzymes. Mammals possess four types of Y-family DNA polymerases: Pol η, Pol κ, Pol ι, and REV1. Among those, Pol η, Pol κ, and Pol ι interact with PCNA through PCNA-binding motifs, low-affinity variants of PCNA-interacting protein box (PIP-box). To date, several studies have reported that REV1 interacts with PCNA, but identified PCNA-binding regions are inconsistent; therefore, a structural basis for interaction between REV1 and PCNA also remains unclear. Here, we identified a signature sequence conserved within vertebrates REV1 responsible for PCNA-binding. Furthermore, we unveiled a mechanism underlying the physical interaction between the PCNA-binding motif of human REV1 and PCNA by X-ray crystallography, thus revealing that REV1 binds to PCNA through a PIP-box variant located in the C-terminal side of the little finger domain. Our study provides a convincing answer for a long-standing controversy regarding the physical interaction between REV1 and PCNA.
- School of Pharmaceutical Sciences, University of Shizuoka, 52-1 Yada, Suruga-ku Shizuoka, 422-8002, Japan.
Organizational Affiliation: