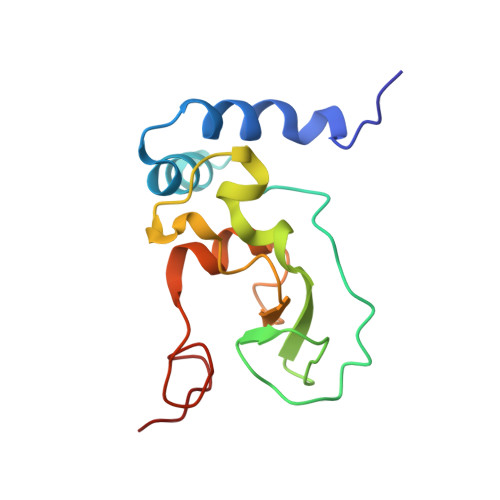
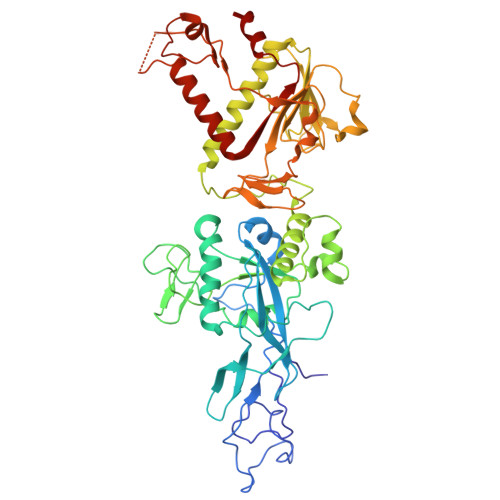
Structural Basis and Rational Design of Nucleotide Analogue Inhibitor Evading the SARS-CoV-2 Proofreading Enzyme.
Wang, J., Pan, Y., Liu, Y., Huang, B., Jin, G., Zhang, L., Zhou, F., Chang, X., Huang, Y., Yan, L., Dong, Y., Rao, Z., Liu, D., Lou, Z.(2025) J Am Chem Soc 147: 21896-21906
- PubMed: 40509573 Search on PubMed
- DOI: https://doi.org/10.1021/jacs.5c05014
- Primary Citation Related Structures:
9VCK, 9VCL - PubMed Abstract:
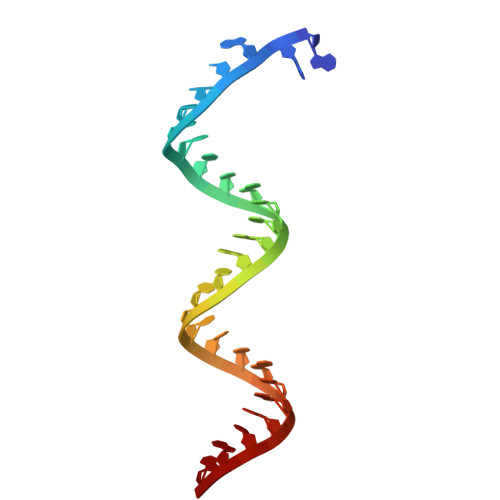
All coronaviruses (CoVs) encode an exoribonuclease in nonstructural protein nsp14 (nsp14 ExoN), which is required for the excision of mismatched nucleotides or nucleotide analogues (NAs) that are incorporated into nascent RNA. Here, we investigated the mechanism by which NAs evade SARS-CoV-2 nsp14 ExoN cleavage using chemically synthesized RNA with NAs incorporated at the 3' end. Nsp14 ExoN exhibited significantly attenuated activity on RNA with sofosbuvir monophosphate (SMP) compared with natural nucleotides, remdesivir/molnupiravir monophosphate, and, in particular, AT-9010 monophosphate (ATMP), which has the same chemically modified ribose moiety as SMP, incorporated at the 3' end. Cryo-electron microscopy structures of nsp10/14 bound to RNA-SMP/-ATMP and mutagenesis studies revealed the essential roles of H95/Q145/F146 in recognizing the base moiety and thus pulling the NAs into a favored conformation for cleavage. Therefore, NAs may evade nsp14 ExoN cleavage by having (1) a base that does not interact with H95, Q145, or F146 and (2) a chemically modified ribose. Guided by this hypothesis, two NAs were designed to effectively resist nsp14 ExoN cleavage. These results inform the rational design of anti-CoV NAs.
- MOE Key Laboratory of Protein Science, School of Medicine, Tsinghua University, Beijing 100084, China.
Organizational Affiliation: