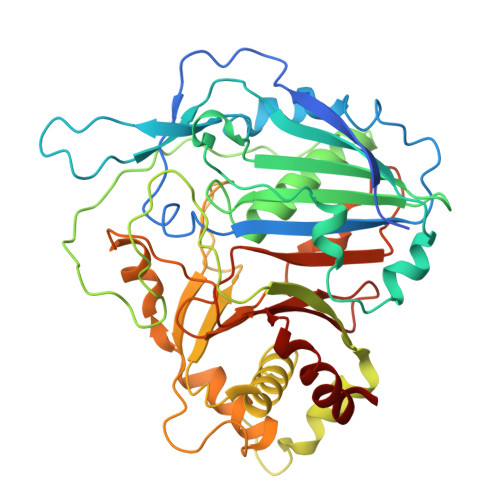
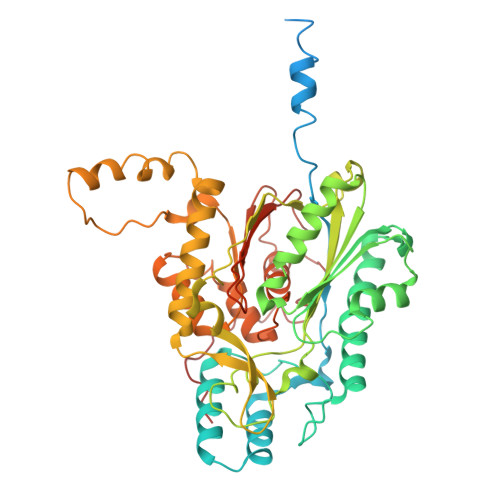
Molecular basis of very-long-chain fatty acid elongation by the CER6-GL2 enzyme complex in plant wax biosynthesis.
Liu, Y., Chen, Y., Zhang, X., Li, M., Wang, J., Yang, Z., Ma, M., Zhao, Z., Liu, H., Yu, F., Zhang, P.(2025) Sci Adv 11: eadz0135-eadz0135
- PubMed: 41337596
- DOI: https://doi.org/10.1126/sciadv.adz0135
- Primary Citation Related Structures:
9UU3, 9UU4, 9UU5 - PubMed Abstract:
Plant cuticular waxes, crucial hydrophobic barriers, are primarily composed of aliphatics derived from very-long-chain fatty acids (VLCFAs; >C28) synthesized by the endoplasmic reticulum fatty acid elongase complex. The core catalytic subunit, CER6 (KCS6), requires interaction with the BAHD protein GL2 to elongate acyl chains beyond C28. We determined the cryo-electron microscopy structure of the maize CER6-GL2 (ZmCER6-ZmGL2) heterotetramer bound to coenzyme A (CoA) and malonyl-CoA, revealing a membrane-anchored ZmCER6 homodimer, with each cytosolic catalytic domain having a substrate tunnel. Structural and biochemical analyses suggest that ZmGL2's amino terminus binds ZmCER6 and remodels its substrate tunnel into a continuous hydrophobic channel at their interface, enabling acyl-chain elongation. CER6 uses a distinct Cys-His-Asn catalytic triad, differing from the histidine-dependent catalysis of mammalian elongases. GL2 acts noncatalytically to modulate CER6 activity. Comparative analyses suggest that species-specific substrate preferences arise from divergent CER2/GL2 interactions. This work elucidates the acyl-chain elongation mechanism of plant VLCFA biosynthesis and provides a foundation for engineering stress-resilient crops via wax modulation.
- Key Laboratory of Plant Carbon Capture, CAS Center for Excellence in Molecular Plant Sciences, Institute of Plant Physiology and Ecology, Chinese Academy of Sciences, Shanghai, China.
Organizational Affiliation: