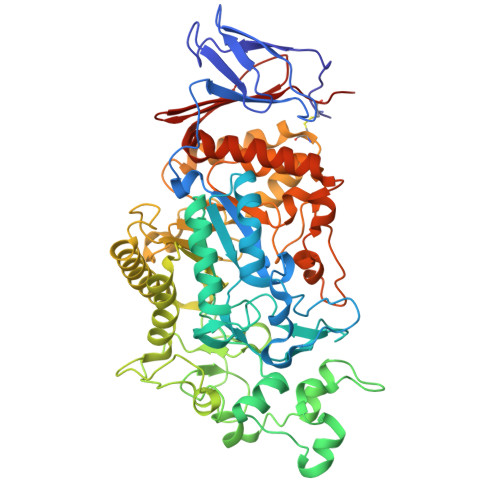
Crystal structure of Klebsiella pneumoniae maltohexaose-producing alpha-amylase.
Fujimoto, Z., Kishine, N., Momma, M.(2025) J Biochem 178: 201-207
- PubMed: 40576559 Search on PubMed
- DOI: https://doi.org/10.1093/jb/mvaf034
- Primary Citation Related Structures:
9US3, 9US4, 9US5, 9US6 - PubMed Abstract:
The α-amylase from Klebsiella pneumoniae (KpAmy13), which belongs to glycoside hydrolase family 13 subfamily 19, produces maltohexaose as an initial product when acting on starch and has been characterized as a maltohexaose-producing α-amylase. The crystal structure of KpAmy13 was determined at a resolution of 1.9 Å, revealing the structures of all its domains: N, A, B, and C. Domain N resembles the starch-binding domain known as carbohydrate-binding module family 69, found in α-glucan-related proteins. Although domain N does not conserve the starch-binding residues observed in other proteins, it has several hydrophobic residues on its surface, which might be involved in promoting catalysis. The catalytic cleft is located at the bottom of a circular depression. The domain N-truncated mutant of KpAmy13 in complex with maltohexaose showed that its non-reducing end glucose docks at subsite -6. The long and complex structure of domain B contributes to forming a cleft of the right size for six glucose moieties, demonstrating the exo-acting mechanism.
- Research Center for Advanced Analysis, National Agriculture and Food Research Organization, 2-1-2 Kannondai, Tsukuba, Ibaraki 305-8518, Japan.
Organizational Affiliation: