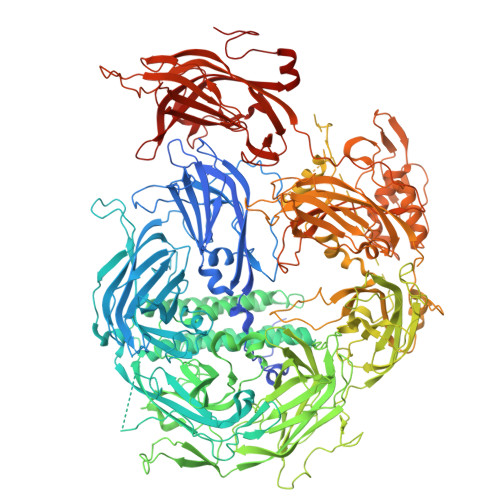
Structure and function of otoferlin, a synaptic protein of sensory hair cells essential for hearing.
Chen, H., Cretu, C., Trebilcock, A., Evdokimova, N., Babai, N., Feldmann, L., Leidner, F., Benseler, F., Mutschall, S., Esch, K., Szabo, C.Z.K., Pena, V., Pape, C., Grubmuller, H., Strenzke, N., Brose, N., Wichmann, C., Preobraschenski, J., Moser, T.(2025) Sci Adv 11: eady8532-eady8532
- PubMed: 41091875 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.ady8532
- Primary Citation Related Structures:
9QE2, 9SE5, 9SEA, 9SEG, 9SFL, 9SH0, 9SI1 - PubMed Abstract:
Hearing relies upon speedy synaptic transmission of sound information from inner hair cells (IHCs) to spiral ganglion neurons. To accomplish this, IHCs use a sophisticated presynaptic machinery including the multi-C 2 domain protein otoferlin that is affected by human deafness mutations. Otoferlin is essential for IHC exocytosis, but how it binds Ca 2+ and the target membrane to serve synaptic vesicle (SV) tethering, docking, and fusion remained unclear. Here, we obtained cryo-electron microscopy structures of otoferlin and employed molecular dynamics simulations of membrane binding. We show that membrane binding by otoferlin involves C 2 B-C 2 G domains and repositions C 2 F and C 2 G domains. Disruption of Ca 2+ -binding sites of the C 2 D domain in mice altered synaptic sound encoding and eliminated the Ca 2+ cooperativity of IHC exocytosis, indicating that it requires the binding of several Ca 2+ -ions by otoferlin. Together, our findings elucidate molecular mechanisms underlying otoferlin-mediated SV docking and support the role of otoferlin as Ca 2+ sensor of SV fusion in IHCs.
- Institute for Auditory Neuroscience and InnerEarLab, University Medical Center Göttingen, Göttingen, Germany.
Organizational Affiliation: