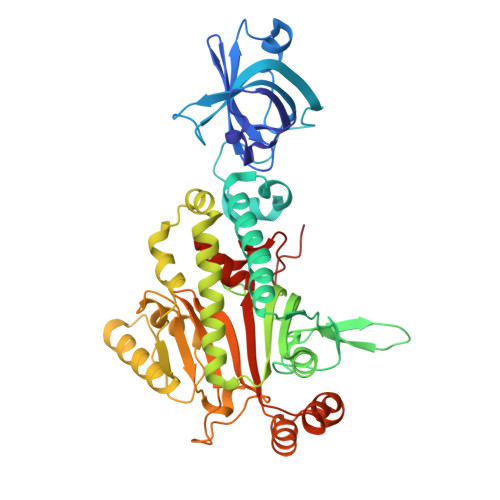
The crystal structure of asparaginyl-tRNA synthetase from Thermus thermophilus and its complexes with ATP and asparaginyl-adenylate: the mechanism of discrimination between asparagine and aspartic acid.
Berthet-Colominas, C., Seignovert, L., Haertlein, M., Grotli, M., Cusack, S., Leberman, R.(1998) EMBO J 17: 2947-2960
- PubMed: 9582288 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/17.10.2947
- Primary Citation Related Structures:
9SH8, 9SH9 - PubMed Abstract:
The crystal structure of Thermus thermophilus asparaginyl-tRNA synthetase has been solved by multiple isomorphous replacement and refined at 2.6 A resolution. This is the last of the three class IIb aminoacyl-tRNA synthetase structures to be determined. As expected from primary sequence comparisons, there are remarkable similarities between the tertiary structures of asparaginyl-tRNA synthetase and aspartyl-tRNA synthetase, and most of the active site residues are identical except for three key differences. The structure at 2.65 A of asparaginyl-tRNA synthetase complexed with a non-hydrolysable analogue of asparaginyl-adenylate permits a detailed explanation of how these three differences allow each enzyme to discriminate between their respective and very similar amino acid substrates, asparagine and aspartic acid. In addition, a structure of the complex of asparaginyl-tRNA synthetase with ATP shows exactly the same configuration of three divalent cations as previously observed in the seryl-tRNA synthetase-ATP complex, showing that this a general feature of class II synthetases. The structural similarity of asparaginyl- and aspartyl-tRNA synthetases as well as that of both enzymes to the ammonia-dependent asparagine synthetase suggests that these three enzymes have evolved relatively recently from a common ancestor.
- European Molecular Biology Laboratory, Grenoble Outstation, Grenoble, France.
Organizational Affiliation: