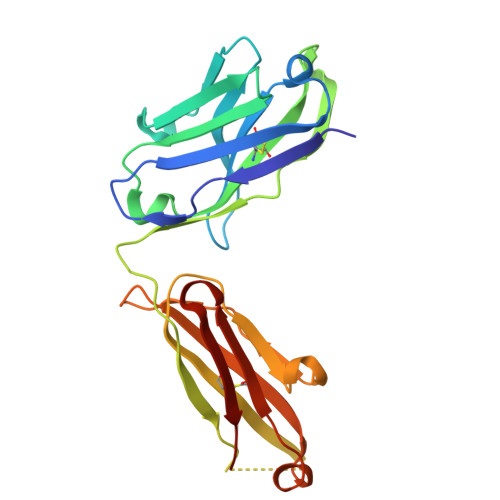
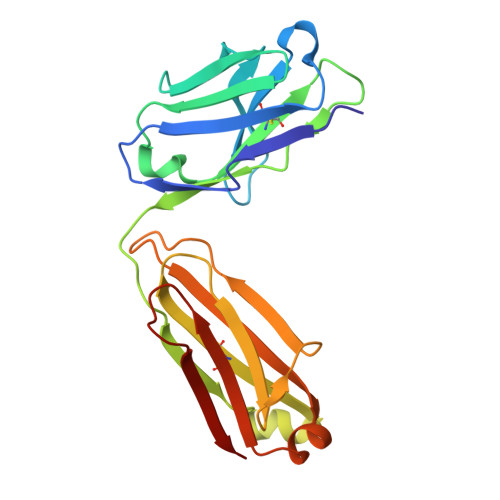
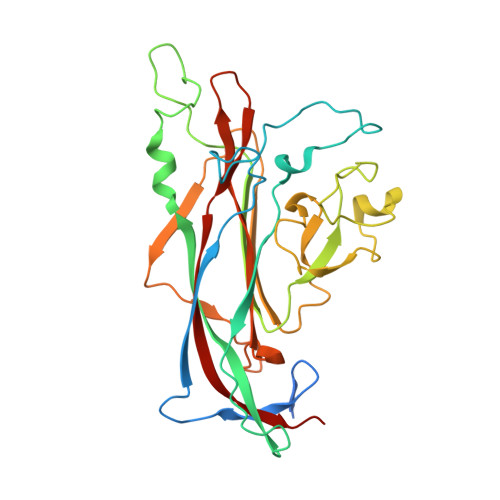
A highly potent human antibody neutralizing all serotypes of BK polyomavirus.
Weber, M., Schmitt, S., Eicher, B., Seidenberg, J., Rutkauskaite, J., Stockli, B., Townsend, C., Huynh-Do, U., Schachtner, T., Delbue, S., Mader, A., Esslinger, C., Hillenbrand, M.(2025) PLoS Pathog 21: e1013122-e1013122
- PubMed: 40680077 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1013122
- Primary Citation Related Structures:
9RM2 - PubMed Abstract:
BK polyomavirus infection poses a significant risk to kidney transplant recipients. Reactivation of dormant virus in the transplanted kidney, triggered by immunosuppression, can lead to BK polyomavirus-associated nephropathy in up to 10% of transplants, often resulting in loss of graft function or even graft loss. Currently, there is no specific treatment that reliably prevents BK polyomavirus-associated nephropathy or halts its progression. Standard of care relies on reducing immunosuppression, allowing the immune system to gradually control the infection but at the risk of provoking rejection episodes and compromising kidney function. This study describes the discovery and characterization of a highly-potent BK polyomavirus-neutralizing antibody, identified from a kidney transplant recipient who displayed rapid clearance of high-level BKPyV-DNAemia. Antibody mAb 319C07 (USAN potravitug) neutralizes all four BK polyomavirus serotypes and recognizes an epitope critical for viral cell attachment. mAb 319C07 is in clinical development for the treatment of BKPyV infection in renal transplant patients (NCT05769582).
- Memo Therapeutics AG, Schlieren Zurich, Zurich, Switzerland.
Organizational Affiliation: