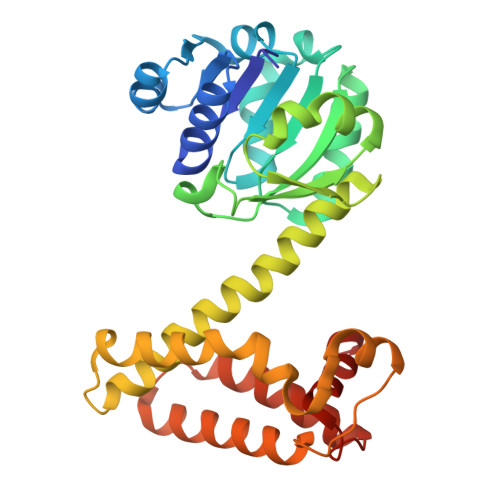
Structures of "Tyrosine-IRED" IR91 from Kribbella flavida in Complex with a Reductive Amination Substrate and Product.
Srinivas, K., Gilio, A.K., Sharma, M., Green, L., Ascham, A., Domenech, J., Pogranyi, B., Li, J., France, S.P., Lewis, R.D., Unsworth, W.P., Grogan, G.(2025) Chembiochem 26: e202500450-e202500450
- PubMed: 40727968 Search on PubMed
- DOI: https://doi.org/10.1002/cbic.202500450
- Primary Citation Related Structures:
9R76, 9R79, 9R7A - PubMed Abstract:
Imine reductases with an (S)-preference for the reduction of the model substrate 2-methyl pyrroline typically contain tyrosine in the active site (Y-IREDs) instead of the aspartate present within (R)-selective enzymes (D-IREDs). As with D-IREDs, a subset of Y-IREDs is capable of enabling reductive amination reactions between some ketone and amine partners to give optically active amines with high optical purity. However, structures of Y-IREDs in complex with the substrates and products of the reductive amination have not been forthcoming. Herein, structures of the Y-IRED IR91 from Kribbella flavida in complex with 5-methoxy-2-tetralone, a synthetic precursor to the anti-Parkinson's treatment rotigotine, and also its reductive amination product with methylamine, 5-methoxy-(S)-2-(N-methylamino)-tetralin, are presented. The structures, in combination with mutation and kinetic studies, support a role for tryptophan residue W258 in the activity of the enzyme, possibly in binding of the ketone prior to reaction with methylamine.
- Department of Chemistry, University of York, Heslington, York, YO10 5DD, UK.
Organizational Affiliation: