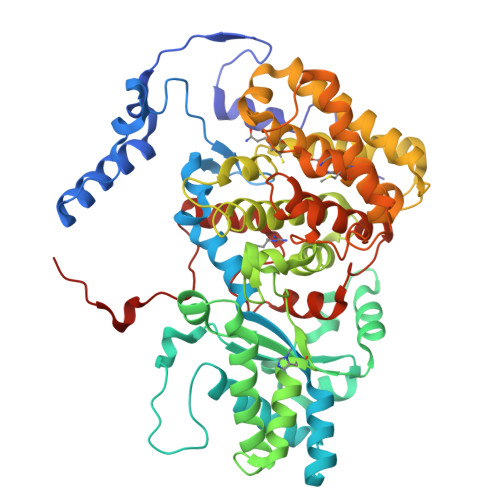
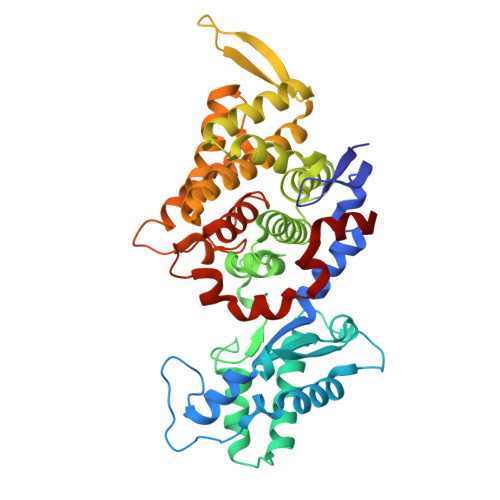
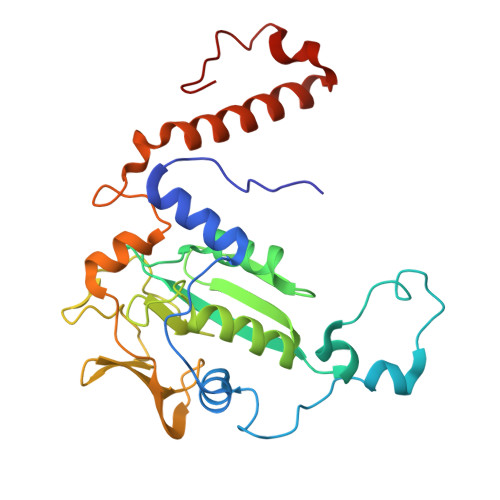
Atomic resolution structures of the methane-activating enzyme in anaerobic methanotrophy reveal extensive post-translational modifications.
Muller, M.C., Wissink, M., Mukherjee, P., Von Possel, N., Laso-Perez, R., Engilberge, S., Carpentier, P., Kahnt, J., Wegener, G., Welte, C.U., Wagner, T.(2025) Nat Commun 16: 8229-8229
- PubMed: 40913044 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-63387-1
- Primary Citation Related Structures:
9QM5, 9QQT, 9QR1, 9QR3 - PubMed Abstract:
Anaerobic methanotrophic archaea (ANME) are crucial to planetary carbon cycling. They oxidise methane in anoxic niches by transferring electrons to nitrate, metal oxides, or sulfate-reducing bacteria. No ANMEs have been isolated, hampering the biochemical investigation of anaerobic methane oxidation. Here, we obtained the true atomic resolution structure of their methane-capturing system (Methyl-Coenzyme M Reductase, MCR), circumventing the isolation barrier by exploiting microbial enrichments of freshwater nitrate-reducing ANME-2d grown in bioreactors, and marine ANME-2c in syntrophy with bacterial partners. Despite their physiological differences, these ANMEs have extremely conserved MCR structures, similar to homologs from methanogenic Methanosarcinales, rather than the phylogenetically distant MCR of ANME-1 isolated from Black Sea mats. The three studied enzymes have seven post-translational modifications, among them was a novel 3(S)-methylhistidine on the γ-chain of both ANME-2d MCRs. Labelling with gaseous krypton did not reveal any internal channels that would facilitate alkane diffusion to the active site, as observed in the ethane-specialised enzyme. Based on our data, the methanotrophic MCRs should follow the same radical reaction mechanism proposed for the methane-generating homologues. The described pattern of post-translational modifications underscores the importance of native purification as a powerful approach to discovering intrinsic enzymatic features in non-isolated microorganisms existing in nature.
- Max-Planck-Institute for Marine Microbiology, Bremen, Germany.
Organizational Affiliation: