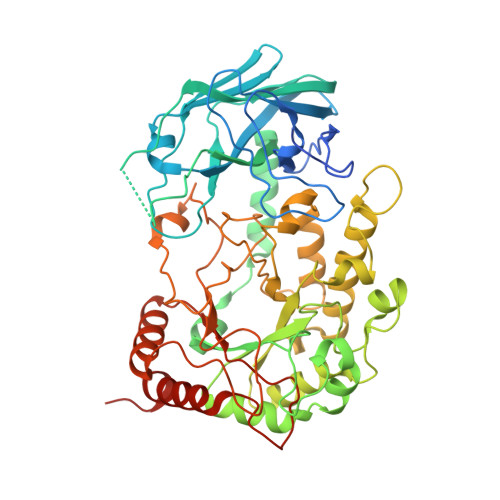
Structural insights into a fucosidase involved in fucoidan degradation.
Bailey, B., Winchester, A., McClain, D., Clingaman, M., Higgins, M.A.(2025) Acta Crystallogr F Struct Biol Commun 81: 459-466
- PubMed: 41114657 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X25008842
- Primary Citation Related Structures:
9PNJ, 9PNK - PubMed Abstract:
Fucoidan is a complex, sulfated polysaccharide primarily found in brown algae, where it plays important structural and protective roles. Due to its abundance in marine ecosystems, many marine bacteria have evolved diverse and specialized enzymatic systems to degrade fucoidan, although the functions and structures of many of these enzymes remain uncharacterized. Here, we describe the structure of a newly identified fucosidase, FucWf4, which cleaves terminal, unsulfated fucose residues from linear, sulfated fucoidan. FucWf4 does not belong to any known glycoside hydrolase (GH) family, but shows the greatest similarity to GH29 fucosidases. We present the first crystal structure of FucWf4 in complex with fucose, revealing a unique C-terminal domain that resembles a carbohydrate-binding module, although it may have lost its carbohydrate-binding capacity and is absent from canonical GH29 enzymes. Docking experiments suggest the presence of a -1 subsite containing a potential sulfate-binding pocket, which may underlie the substrate specificity of the enzyme. Furthermore, sequence analysis of FucWf4 homologs reveals two distinct clades, likely corresponding to functionally divergent groups. Together, these findings provide new insights into the molecular basis of fucoidan recognition and degradation by this novel enzyme subfamily, laying the groundwork for future functional and structural studies.
- Department of Biological Sciences, University of Alabama, Tuscaloosa, 3314 Science and Engineering Complex, Tuscaloosa, AL 35487, USA.
Organizational Affiliation: