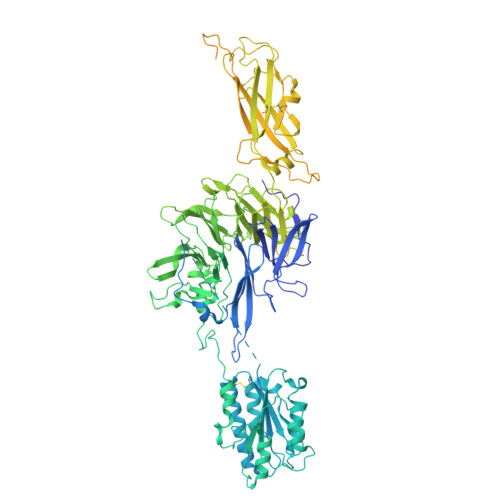
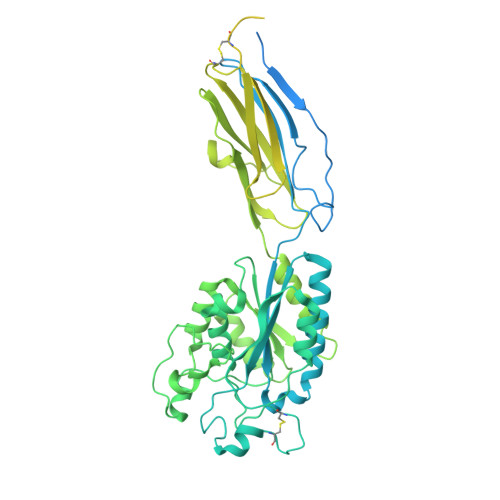
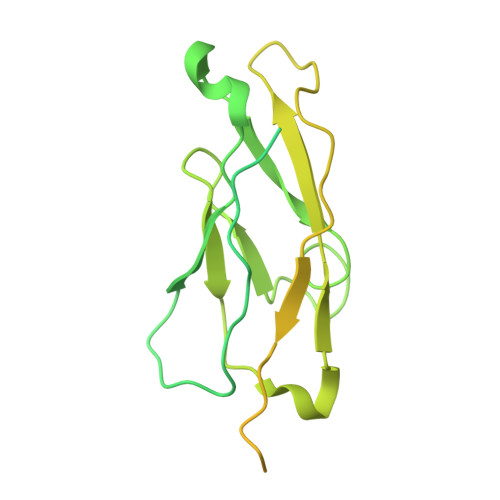
Molecular exaptation by the integrin alpha I domain.
Hollis, J.A., Chan, M.C., Malik, H.S., Campbell, M.G.(2025) Sci Adv 11: eadx9567-eadx9567
- PubMed: 40929264 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.adx9567
- Primary Citation Related Structures:
9P95, 9P96, 9P97, 9P98, 9P99 - PubMed Abstract:
Integrins bind ligands between their alpha (α) and beta (β) subunits and transmit signals through conformational changes. Early in chordate evolution, some α subunits acquired an "inserted" (I) domain that expanded integrin's ligand-binding repertoire but obstructed the ancestral ligand pocket, seemingly blocking conventional integrin activation. Here, we compare cryo-electron microscopy structures of apo and ligand-bound states of the I domain-containing αEβ 7 integrin and the I domain-lacking α 4 β 7 integrin to illuminate how the I domain intrinsically mimics an extrinsic ligand to preserve integrin function. We trace the I domain's evolutionary origin to an ancestral collagen-collagen interaction domain, identifying an ancient molecular exaptation that facilitated integrin activation immediately upon I domain insertion. Our analyses reveal the evolutionary and biochemical basis of expanded cellular communication in vertebrates.
- Division of Basic Sciences, Fred Hutchinson Cancer Center, Seattle, WA 98109, USA.
Organizational Affiliation: