RO4, a high-affinity humanized antibody against the juxtamembrane region of mesothelin for targeted cancer therapy.
Onda, M., Liu, X., Liu, W., Zhan, J., Maslanka, C.A., Xia, D., Ho, M., Pastan, I.(2026) Antib Ther 9: 1-12
- PubMed: 41550974
- DOI: https://doi.org/10.1093/abt/tbaf022
- Primary Citation of Related Structures:
9P4C - PubMed Abstract:
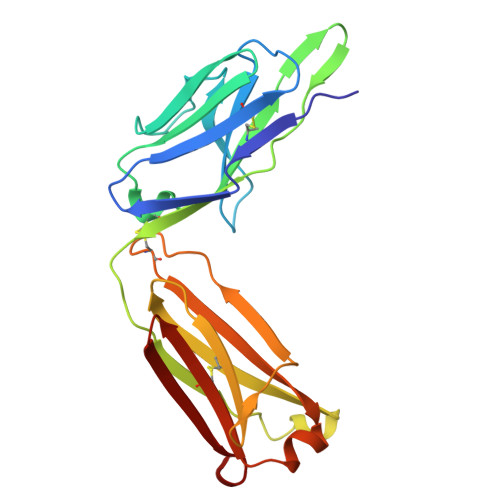
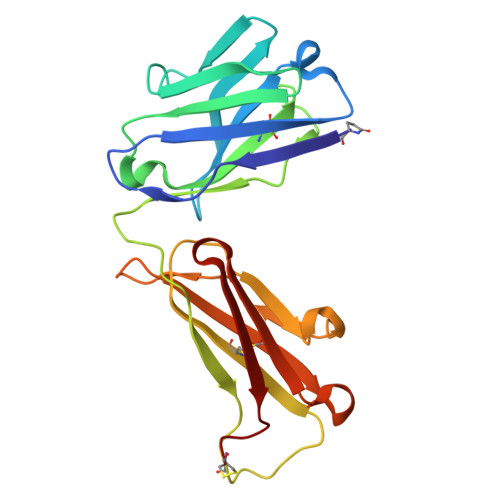
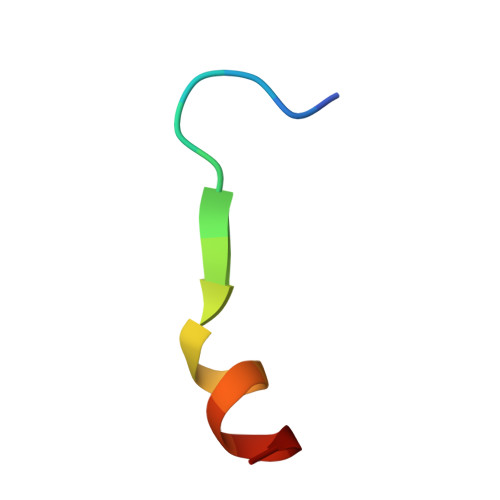
Mesothelin (MSLN) is a surface antigen highly expressed in several solid tumors, including mesothelioma, ovarian, and pancreatic cancers. However, therapeutic efficacy of MSLN-targeted agents is often compromised by shed MSLN (SM), which acts as a soluble decoy and accumulates in tumor microenvironments, reducing antibody engagement at the tumor surface. To overcome this barrier, we generated antibodies targeting the membrane-proximal, non-shed region of MSLN using a peptide encompassing major cleavage sites for rabbit immunization. From 200 B-cell clones, 14 antibodies specific to the juxtamembrane region of MSLN were identified. The lead candidate, RO4, underwent detailed characterization and humanization to improve clinical applicability. Humanized RO4 (hRO4) exhibited enhanced binding affinity to MSLN and specifically recognized tumor-associated, non-shed epitopes. Structural analysis confirmed precise epitope engagement near the cleavage site. hRO4 effectively inhibited mesothelin shedding in vitro and enabled potent tumor eradication when expressed in chimeric antigen receptor (CAR) T cells in NOD scid gamma mouse models. Targeting a non-shed epitope of MSLN with hRO4 overcomes a critical limitation of conventional MSLN-directed therapies. By avoiding decoy interference and enhancing tumor-specific targeting, hRO4-based therapeutics offer promising clinical potential for improving outcomes in MSLN-positive cancers.
- Laboratory of Molecular Biology, Center for Cancer Research, National Cancer Institute, National Institutes of Health, Bethesda, MD 20892, United States.
Organizational Affiliation: