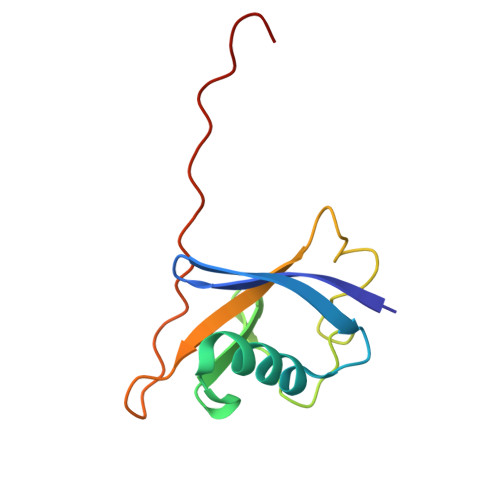
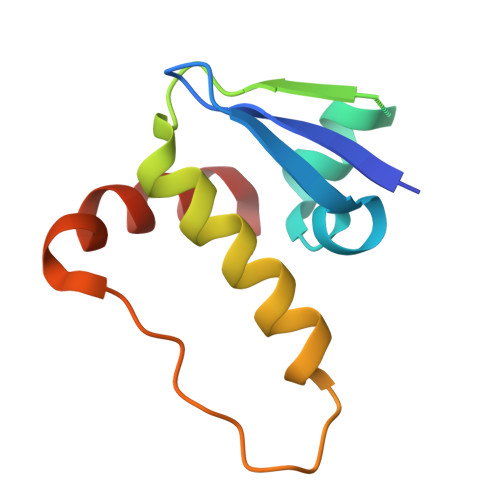
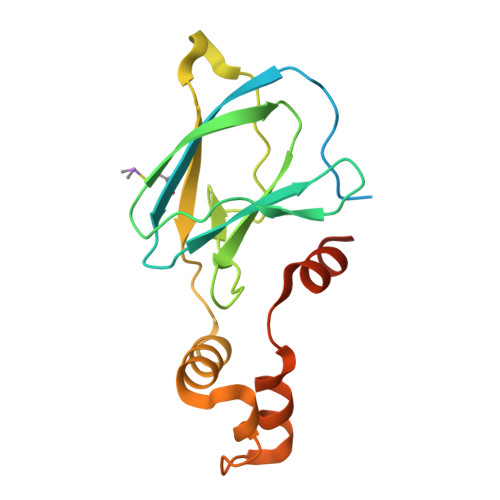
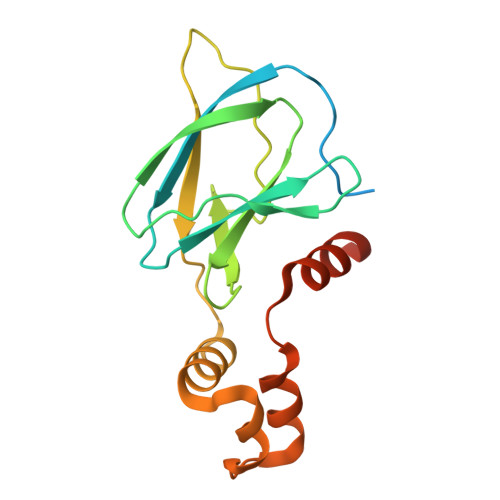
NMR-Based Fragment Screen of the von Hippel-Lindau Elongin C&B Complex.
Amporndanai, K., Katinas, J.M., Chopra, A., Kayode, O., Vadukoot, A.K., Waterson, A.G., Fesik, S.W.(2025) ACS Med Chem Lett 16: 1648-1654
- PubMed: 40832550 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsmedchemlett.5c00316
- Primary Citation Related Structures:
9OIM, 9OIN, 9OIO, 9OIQ - PubMed Abstract:
von Hippel-Lindau (VHL) is an E3 ligase that has been widely exploited for the development of PROTACs to induce degradation of disease-associated target proteins. Nearly all VHL-recruiting PROTACs contain a hydroxyproline moiety based on the endogenous peptide substrate that occupies the HIF1α-binding site of VHL. However, the development of orally bioavailable PROTACs with hydroxyproline-based VHL ligands remains a significant hurdle, due to both the hydroxyproline and the peptidic nature of the VHL ligand. Here, we describe an NMR-based fragment screen against the VHL-Elongin C-Elongin B (VCB) complex. Several hits were shown by X-ray crystallography to bind to the HIF1α active site in VHL of the VCB complex, which opens the possibility for the discovery of new nonhydroxyproline-based VHL ligands for use in VHL-recruiting PROTACs.
- Department of Biochemistry, Vanderbilt University School of Medicine, Nashville, Tennessee 37232-0146, United States.
Organizational Affiliation: