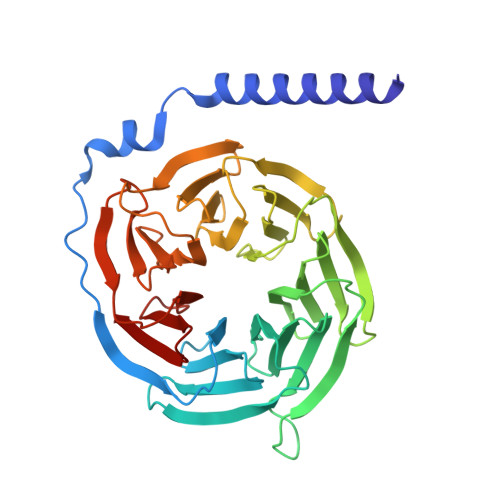
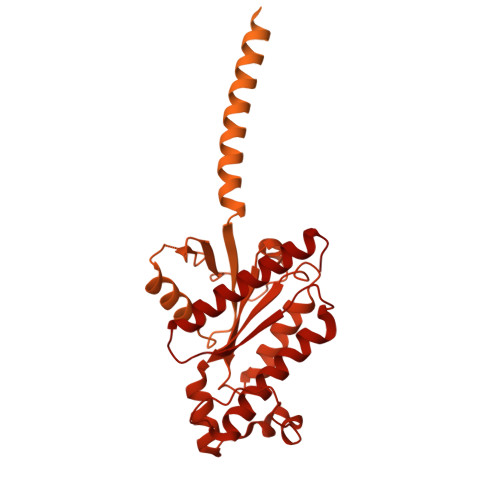
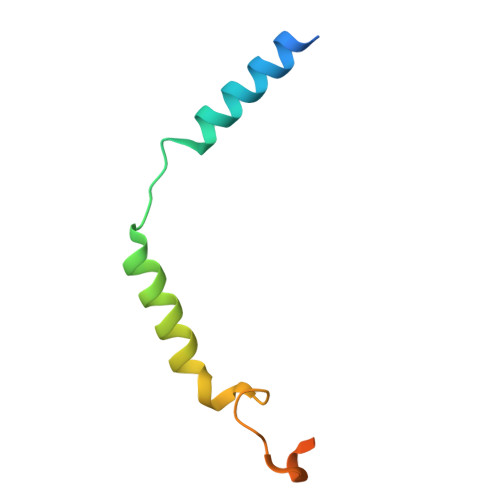
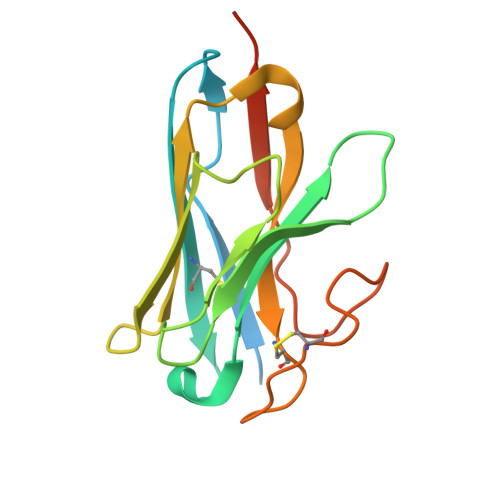
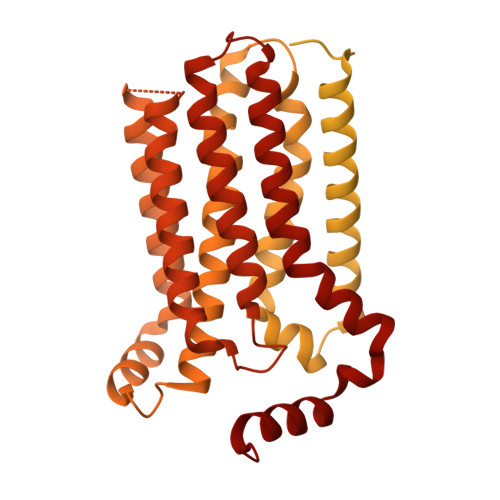
The structure of human sweetness.
Juen, Z., Lu, Z., Yu, R., Chang, A.N., Wang, B., Fitzpatrick, A.W.P., Zuker, C.S.(2025) Cell 188: 4141-4153.e18
- PubMed: 40339580 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2025.04.021
- Primary Citation Related Structures:
9NOR, 9NOS, 9NOT, 9NOU, 9NOV, 9NOW, 9NOX, 9NOY, 9O38 - PubMed Abstract:
In humans, the detection and ultimately the perception of sweetness begin in the oral cavity, where taste receptor cells (TRCs) dedicated to sweet-sensing interact with sugars, artificial sweeteners, and other sweet-tasting chemicals. Human sweet TRCs express on their cell surface a sweet receptor that initiates the cascade of signaling events responsible for our strong attraction to sweet stimuli. Here, we describe the cryo-electron microscopy (cryo-EM) structure of the human sweet receptor bound to two of the most widely used artificial sweeteners-sucralose and aspartame. Our results reveal the structural basis for sweet detection, provide insights into how a single receptor mediates all our responses to such a wide range of sweet-tasting compounds, and open up unique possibilities for designing a generation of taste modulators informed by the structure of the human receptor.
- Zuckerman Mind Brain Behavior Institute and Department of Biochemistry and Molecular Biophysics, Vagelos College of Physicians and Surgeons, Columbia University, New York, NY, USA; Howard Hughes Medical Institute, Vagelos College of Physicians and Surgeons, Columbia University, New York, NY 10032, USA.
Organizational Affiliation: