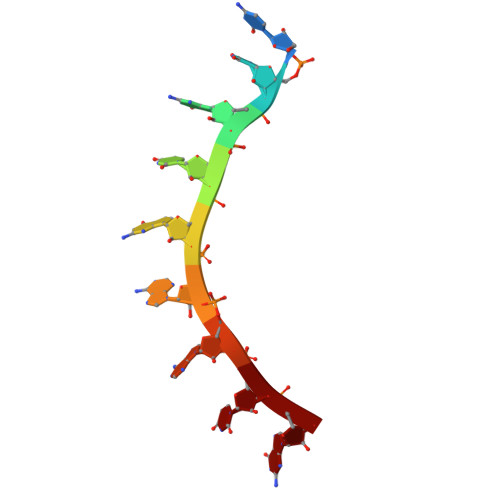
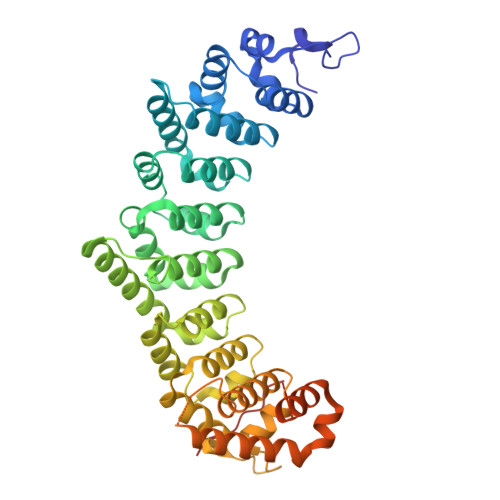
Caenorhabditis elegans FBF-1 and FBF-2 C-terminal intrinsically disordered regions differentially regulate RNA-binding affinity.
Hawthorne, H.R., Qiu, C., Tanaka Hall, T.M.(2025) RNA 31: 1391-1402
- PubMed: 40769718 Search on PubMed
- DOI: https://doi.org/10.1261/rna.080578.125
- Primary Citation Related Structures:
9NM3 - PubMed Abstract:
PUF proteins (named for Drosophila melanogaster Pumilio and Caenorhabditis elegans fem-3 mRNA binding factor or FBF) are a family of RNA-binding proteins. C. elegans FBF is a collective term for two PUF proteins, FBF-1 and FBF-2, that maintain germline stem cells. FBF binds the 3'UTR of target RNAs and together with partner proteins represses translation of mRNAs that promote differentiation. Until recently, little was known about the functions of the FBF C-terminal intrinsically-disordered regions that follow the RNA-binding domain (RBD). Despite high overall protein sequence conservation (91% identical residues), the FBF-1 and FBF-2 C-terminal tails (CTs) are distinct, and the FBF-2 CT is essential for its function. The FBF-2 CT contains a PUF-interacting motif (PIM) that binds its own RBD and autoinhibits RNA-binding affinity. Here we investigated whether differences in the FBF-1 and FBF-2 CTs impact molecular function. Unlike FBF-2, the FBF-1 CT had no impact on RNA binding. Despite this, a crystal structure of FBF-1 demonstrated that a PIM in the FBF-1 CT binds to its RBD, like FBF-2. By creating FBF-1/FBF-2 chimeric proteins, we discovered that the FBF-2 CT can autoinhibit FBF-1 RNA binding, and substitution of the FBF-1 PIM for the FBF-2 PIM diminished FBF-2 autoinhibition. Our results exemplify how RBP paralogs diverge to fine tune their RNA-binding activities.
- NIEHS, NIH.
Organizational Affiliation: