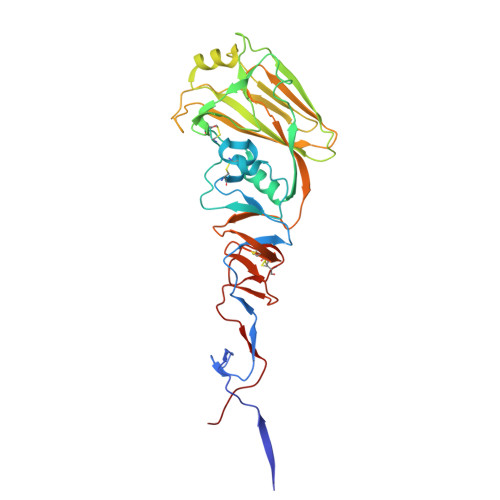
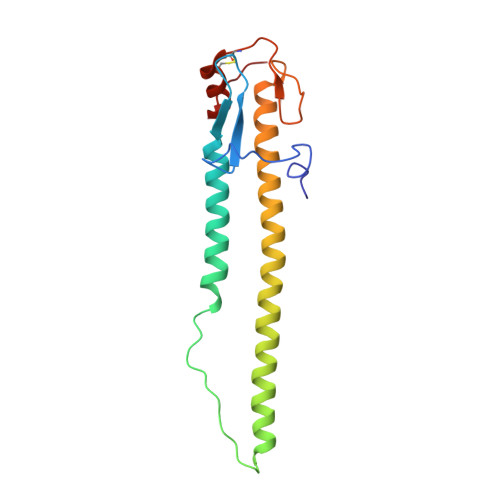
Virion-associated influenza hemagglutinin clusters upon sialic acid binding visualized by cryoelectron tomography.
Huang, Q.J., Kim, R., Song, K., Grigorieff, N., Munro, J.B., Schiffer, C.A., Somasundaran, M.(2025) Proc Natl Acad Sci U S A 122: e2426427122-e2426427122
- PubMed: 40244672 Search on PubMed
- DOI: https://doi.org/10.1073/pnas.2426427122
- Primary Citation Related Structures:
9N8P - PubMed Abstract:
Influenza viruses are enveloped, negative-sense single-stranded RNA viruses covered in a dense layer of glycoproteins. Hemagglutinin (HA) accounts for 80 to 90% of influenza glycoprotein and plays a role in host cell binding and membrane fusion. While previous studies have characterized structures of purified receptor-free and receptor-bound HA, the effect of receptor binding on HA organization and structure on virions remains unknown. Here, we used cryoelectron tomography to visualize influenza virions bound to a sialic acid receptor mimic. Overall, receptor binding did not result in significant changes in viral morphology; however, we observed rearrangements of HA trimer organization and orientation. Compared to the even interglycoprotein spacing of unliganded HA trimers, receptor binding promotes HA trimer clustering and the formation of a triplet of trimers. Subtomogram averaging and refinement yielded 8 to 10 Å reconstructions that allowed us to visualize specific contacts between HAs from neighboring trimers and identify molecular features that mediate clustering. Taken together, we present structural evidence that receptor binding triggers clustering of HA trimers, revealing an additional layer of HA dynamics and plasticity.
- Department of Biochemistry and Molecular Biotechnology, University of Massachusetts Chan Medical School, Worcester, MA 01605.
Organizational Affiliation: