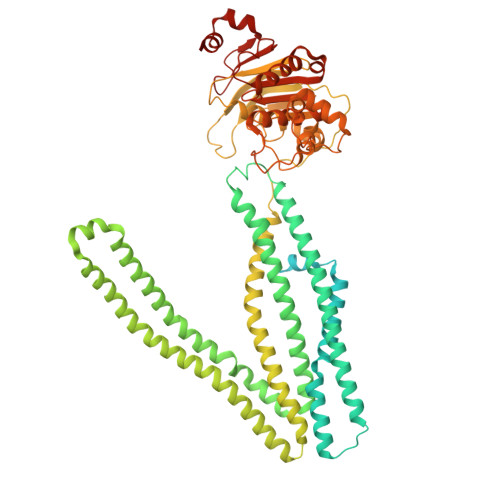
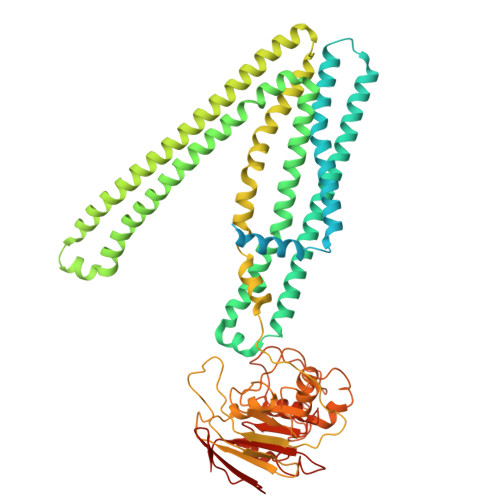
Nucleotide-dependent conformational changes direct peptide export by the transporter associated with antigen processing.
Lee, J., Manon, V., Chen, J.(2025) Immunity 58: 2166-2175.e4
- PubMed: 40885191 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.immuni.2025.08.003
- Primary Citation Related Structures:
9N61, 9N62, 9N63, 9N64, 9N65, 9N66 - PubMed Abstract:
The transporter associated with antigen processing (TAP) delivers peptide antigens from the cytoplasm into the endoplasmic reticulum (ER) for loading onto major histocompatibility complex class I (MHC-I) molecules. To examine the mechanisms of peptide transport and release into the ER, we determined cryo-electron microscopy structures of the human TAP heterodimer in multiple functional states along the transport cycle. In the inward-facing conformation, when the peptide translocation cavity within the TAP heterodimer is exposed to the cytosol, ATP binding strengthened intradomain assembly. Transition to the outward-facing conformation, when the transporter opens to the ER lumen, led to a complete reconfiguration of the peptide-binding site, facilitating peptide release. ATP hydrolysis opened the catalytically active nucleotide-binding consensus site, and the subsequent separation of the nucleotide-binding domains reset the transport cycle. These findings establish a comprehensive structural framework for understanding unilateral peptide transport, vanadate trapping, and trans-inhibition-an internal feedback mechanism that prevents excessive peptide accumulation and activation of the ER stress response.
- Laboratory of Membrane Biophysics and Biology, the Rockefeller University, New York, NY 10065, USA.
Organizational Affiliation: