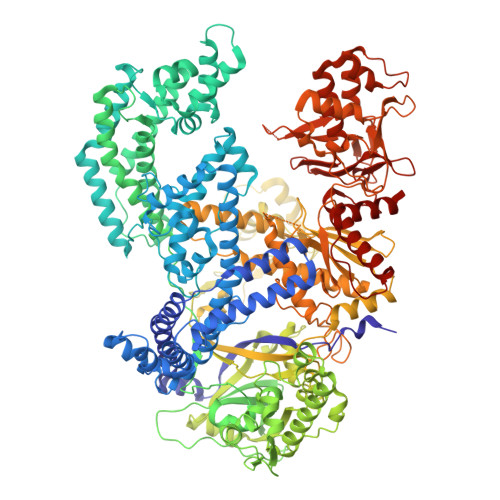
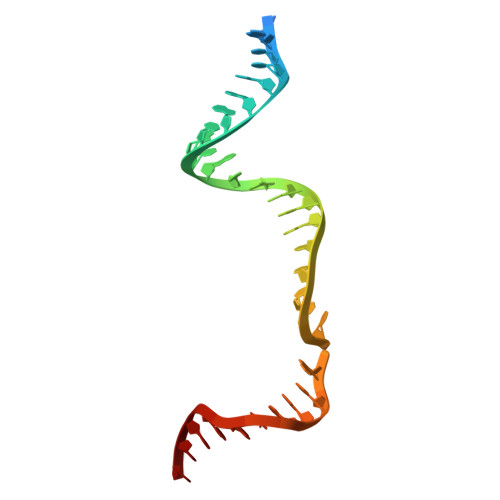
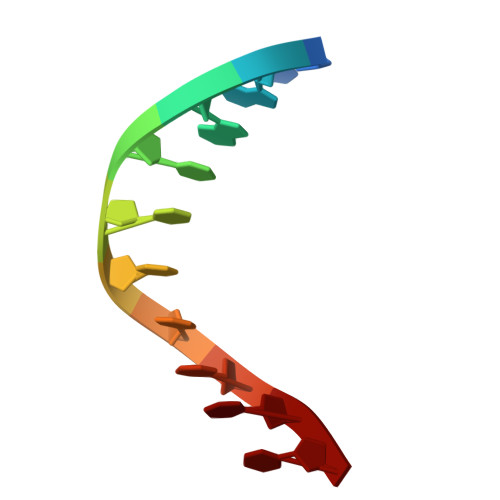
Mechanistic basis for improved activity of Engineered AsCas12a.
Jansson-Fritzberg, L., Chica, B., Latrick, C., Olland, A., Dementiev, A., White, A., Kutter, S., Lemercier, J.N., Wolk, S.(2026) Commun Biol 9
- PubMed: 41807709 Search on PubMed
- DOI: https://doi.org/10.1038/s42003-026-09799-1
- Primary Citation Related Structures:
9MK3 - PubMed Abstract:
CRISPR-associated proteins (Cas) are central to gene editing, forming nuclease complexes with guide RNA to enable precise genome modification. Among numerous Cas variants, Cas9 and Cas12a are the most extensively studied. While much is known about the genomic substrates for these enzymes, less is known about the determinants of the DNA cleavage activity. Wild-type Cas12a exhibits higher intrinsic specificity than Cas9, minimizing off-target activity, but lower overall potency. Recent protein engineering has sought to improve both parameters. Here, we shed light on the structural and mechanistic basis by which an engineered AsCas12a variant achieves high potency while retaining its hallmark specificity. We show that reduced protein-DNA interactions facilitate more rapid R-loop formation, thereby enhancing cleavage activity. These results provide mechanistic insight into Cas12a function and highlight strategies for designing genome-editing nucleases with optimal balance between efficiency and specificity.
- Editas Medicine, Inc, Cambridge, MA, USA.
Organizational Affiliation: