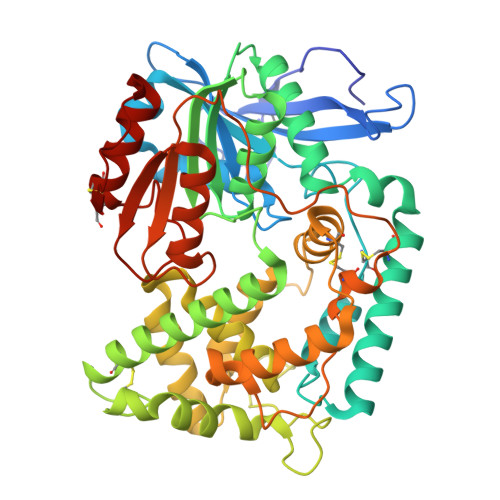
Discovery of highly potent alpha-keto ester-based peptidomimetic inhibitors of the Hip1 protease for the treatment of Mycobacterium tuberculosis.
Schumann, N., Shamma, F., Brooks, C.L., Johnson, S.J., Yim, M.K., Olsen, K.J., Pena, K., Karakousis, P.C., Abell, A., Goldfarb, N.E.(2025) Eur J Med Chem Rep 15
- PubMed: 41929655 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.ejmcr.2025.100311
- Primary Citation Related Structures:
9MD7, 9MD8 - PubMed Abstract:
Mycobacterium tuberculosis (Mtb), the bacterium responsible for tuberculosis, is the leading cause of death due to a single infectious agent. Given the alarming increase in drug-resistant cases, therapeutic agents targeting novel Mtb drug targets are urgently needed. Hip1, a serine protease required for Mtb survival in macrophages and tolerance to various antibiotics, has been identified as an attractive therapeutic target. In the current study, we describe the design and synthesis of highly potent (pM range K i ) peptidomimetic α-keto ester inhibitors of Hip1. We also report the first two X-ray cocrystal structures of Hip1 bound to these compounds and describe the binding interactions in the active site of recombinant Hip1. Finally, we show that these compounds effectively reduce the intracellular bacillary burden in a macrophage model of Mtb infection.
- Department of Chemistry, The University of Adelaide, North Terrace Campus, Adelaide, SA, 5005, Australia.
Organizational Affiliation: