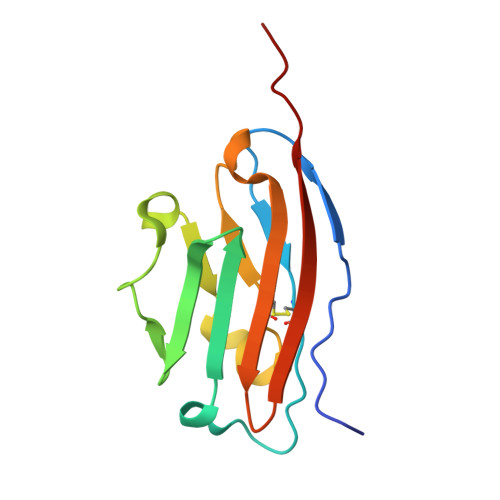
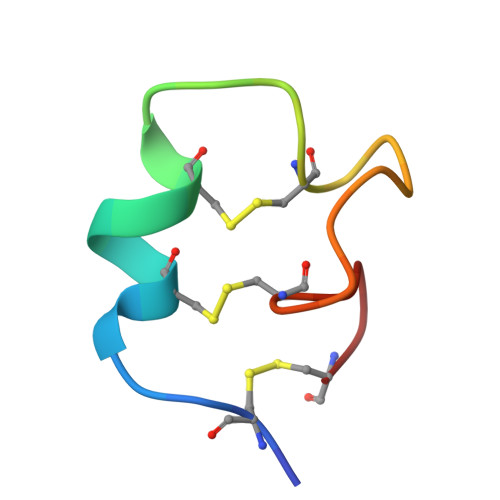
Multicyclic Peptides Targeting PD-L1 for Radiotheranostics: From Discovery to Clinical Proof-of-Concept.
Cheng, X., Jiang, S., Peng, X., Yang, P., Zhang, S., Xu, C., Gao, X., Fan, S., Liu, H., Zhuang, J., Chen, X., Liang, N., Lin, B., Lu, Q., Chen, M., Xiao, Y., Zhu, Z., Wang, R., Hu, K., Wu, C.(2025) J Am Chem Soc 147: 35638-35654
- PubMed: 40974147 Search on PubMed
- DOI: https://doi.org/10.1021/jacs.5c11292
- Primary Citation Related Structures:
9MAP - PubMed Abstract:
Radiotheranostics holds transformative potential for precision oncology by integrating diagnostic imaging with targeted radionuclide therapy. However, advancements in this field are significantly hindered by the limited availability of high-affinity ligands that are capable of engaging challenging cell-surface antigens, particularly flat, low-druggability targets such as programmed death-ligand 1 (PD-L1). Here, we overcome this barrier through de novo discovery and rational engineering of a disulfide-directed multicyclic peptide (DDMP), dmp10, which achieves a picomolar affinity for PD-L1 by leveraging conformationally constrained structural scaffolds. By combining disulfide-directed library design with iterative directed evolution, we successfully generated dmp10, a ∼3 kDa multicyclic peptide that establishes unprecedented shape complementarity to the expansive binding interface of PD-L1. Preclinical evaluations demonstrated that 68 Ga-labeled dmp10 enables high-contrast PET imaging of PD-L1 + tumors in murine models, achieving a tumor uptake of 13.27 %ID/g at 4 h post-injection. The therapeutic counterpart, 177 Lu-labeled dmp10, effectively eradicated 92.47% of established tumors in tumor models while sparing healthy tissues, thereby validating its dual radiotheranostic utility. The translational relevance of our findings was further confirmed in a first-in-human pilot study, where 68 Ga-labeled dmp10 was well tolerated and allowed visualization of PD-L1 + lesions in patients with solid tumors. This work not only establishes DDMPs as a versatile platform for targeting geometrically complex antigens but also delivers a promising radiotheranostic agent that bridges molecular imaging and precision radionuclide therapy for PD-L1-driven malignancies. Our findings advance current strategies for designing ultrahigh-affinity peptide binders and underscore the untapped potential of multicyclic architectures in overcoming longstanding challenges in cancer theranostics.
- Department of Chemistry, College of Chemistry and Chemical Engineering, The MOE Key Laboratory of Spectrochemical Analysis and Instrumentation, State Key Laboratory of Physical Chemistry of Solid Surfaces, Xiamen University, Xiamen 361005, P.R. China.
Organizational Affiliation: