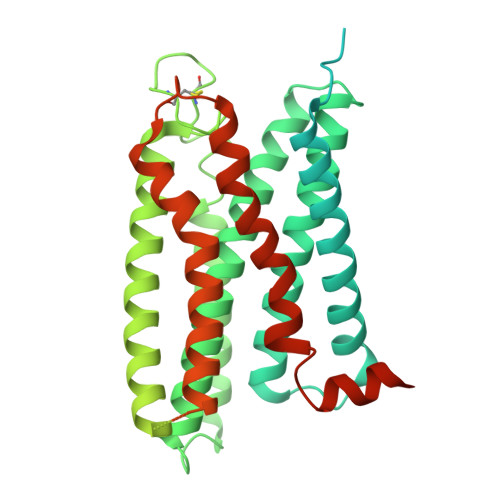
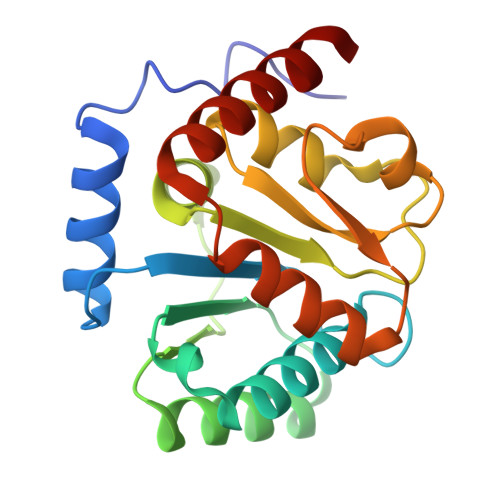
Mechanism and function of GPR3 regulated by a negative allosteric modulator.
Chen, G., Blahova, J., Staffen, N., Hubner, H., Nunhofer, N., Qiu, C., Gmeiner, P., Weikert, D., Du, Y., Xu, J.(2025) Nat Commun 16: 7988-7988
- PubMed: 40866348 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-63422-1
- Primary Citation Related Structures:
9M88, 9M8P, 9M8V - PubMed Abstract:
Allosteric modulators have gained substantial interest in current GPCR drug discovery. Here, we present a mechanism of allosteric modulation involving the dimerization of GPR3, a promising drug target for metabolic diseases and central nervous system disorders. We show that GPR3 forms constitutive homodimers in live cells and reveal that the inhibitor AF64394 functions as a negative allosteric modulator (NAM) specifically targeting dimeric GPR3. Using cryogenic electron microscopy (cryo-EM), we determine the structures of the AF64394-bound GPR3 dimer and its dimer-Gs signaling complex. These high-resolution structures reveal that AF64394 binds to the transmembrane dimer interface. AF64394 binding prevents the dissociation of the GPR3 dimer upon engagement with Gs and restrains transmembrane helix 5 in an inactive-like intermediate conformation, leading to reduced coupling with Gs. Our studies unveil a mechanism of dimer-specific inhibition of signaling with significant implications for the discovery of drugs targeting GPCRs capable of dimerization.
- School of Medicine, Shenzhen Campus of Sun Yat-sen University, Sun Yat-sen University, Shenzhen, China.
Organizational Affiliation: