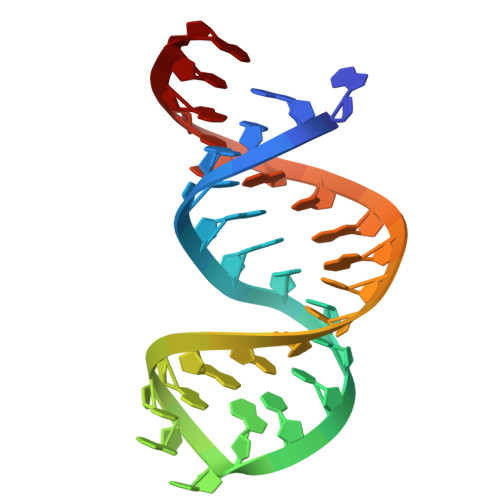
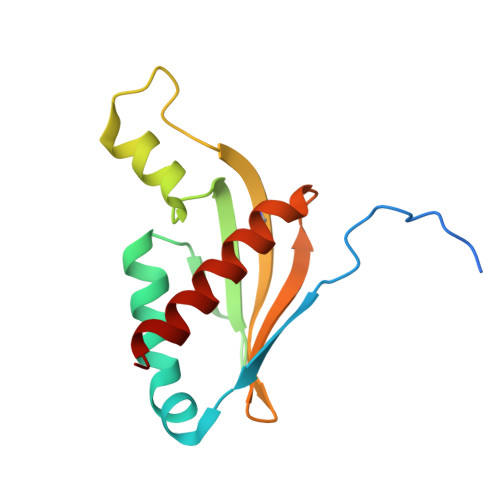
Structures and mechanisms of U6 snRNA m 6 A modification by METTL16.
Ju, J., Tomita, K.(2025) Nat Commun 16: 7708-7708
- PubMed: 40841561 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-63021-0
- Primary Citation Related Structures:
9M86, 9U47, 9U48 - PubMed Abstract:
The N 6 -methyladenosine (m 6 A) modification in U6 snRNA, catalyzed by METTL16 using S-adenosylmethionine (SAM) as the methyl donor, is required for efficient and accurate pre-mRNA splicing. However, the mechanism by which METTL16 modifies U6 snRNA with m 6 A remains elusive. Here, we present cryo-EM structures of METTL16 in complex with U6 snRNA, providing insights into the METTL16-mediated modification of U6 snRNA with m 6 A. The structures reveal that U6 snRNA is recruited to METTL16 through specific interactions between the C-terminal kinase-associated 1 (KA-1) domain of METTL16 and the internal stem-loop (ISL) of U6 snRNA. Upon SAM binding to the catalytic pocket within the N-terminal methyltransferase domain (MTD), U6 snRNA undergoes a structural rearrangement that positions the target adenine-containing motif at the catalytic site. This conformational change is followed by an additional structural adjustment of U6 snRNA into a productive conformation, bringing the target adenosine closer to SAM within the catalytic pocket and thereby ensuring efficient m 6 A modification. The KA-1 domain functions as a scaffold for initial substrate recognition and facilitates the subsequent dynamic methylation process within the MTD, highlighting the cooperative roles of METTL16 domains for U6 snRNA modification.
- Department of Computational Biology and Medical Sciences, Graduate School of Frontier Sciences, The University of Tokyo Kashiwa, Chiba, Japan.
Organizational Affiliation: