Structural basis of auxin binding and transport by Arabidopsis thaliana AUX1.
Jing, D., Kong, F., Lu, X., Huang, G., Huang, J., Wang, H., Shi, Y., Wang, C.(2025) Proc Natl Acad Sci U S A 122: e2513424122-e2513424122
- PubMed: 40720658 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2513424122
- Primary Citation Related Structures:
9LVA, 9LVB - PubMed Abstract:
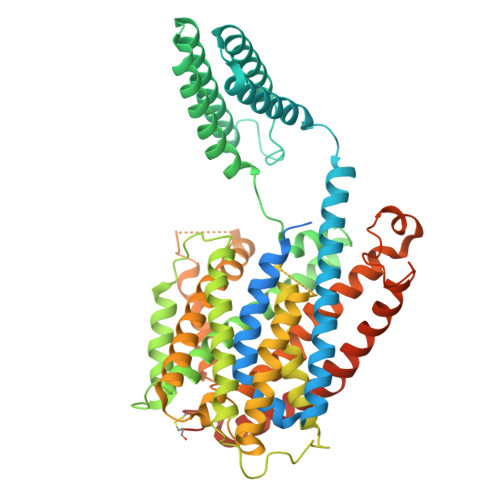
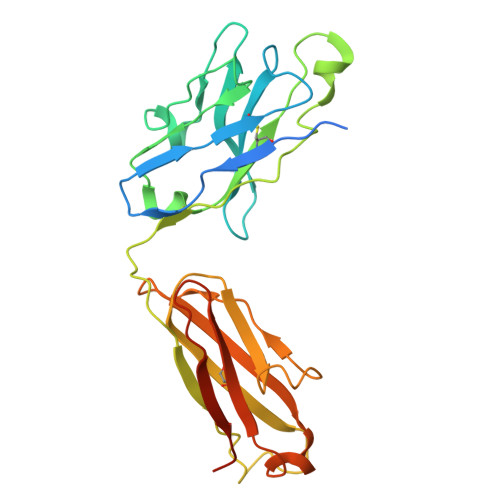
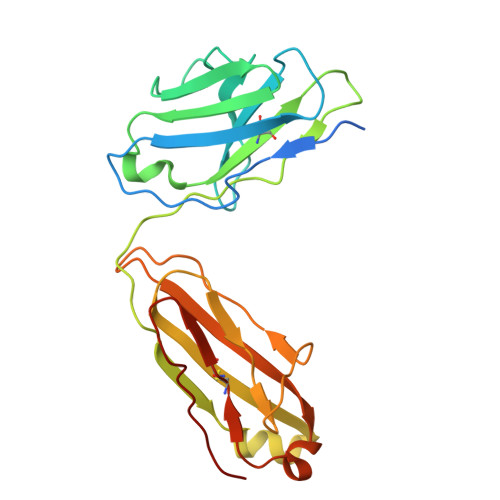
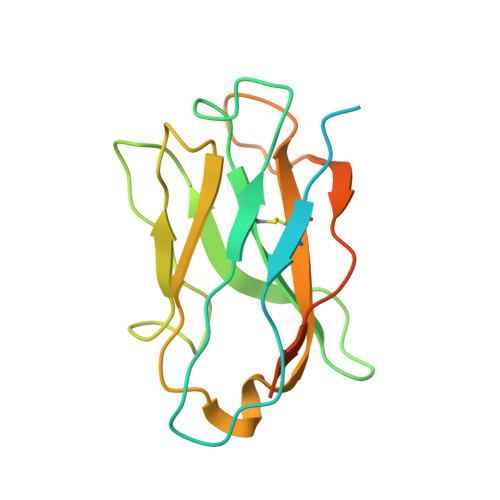
Indole-3-acetic acid (IAA), the major form of auxin, is essential for plant growth. Auxin resistant 1 (AUX1), the first identified auxin importer, plays a crucial role in polar auxin transport (PAT). Here, we present cryo-EM structures of Arabidopsis thaliana AUX1 in the IAA-free and IAA-bound states. AUX1 exists as a monomer that contains 11 transmembrane helices (TMs). TMs 1 to 5 and 6 to 10 constitute the two halves of a classic LeuT-fold, and TM11 interacts with both halves at the interface. In the IAA-bound state, IAA is specifically recognized in a central pocket formed by TM1, TM3, TM6, and TM8. In the presence of IAA, TM1 and TM6 undergo marked conformational changes that are critical for IAA transport. His249 stands out to be a key residue for substrate uptake and release. Our structures reveal the molecular basis for AUX1-mediated IAA binding and transport.
- College of Life Sciences, Zhejiang University, Hangzhou 310058, China.
Organizational Affiliation: