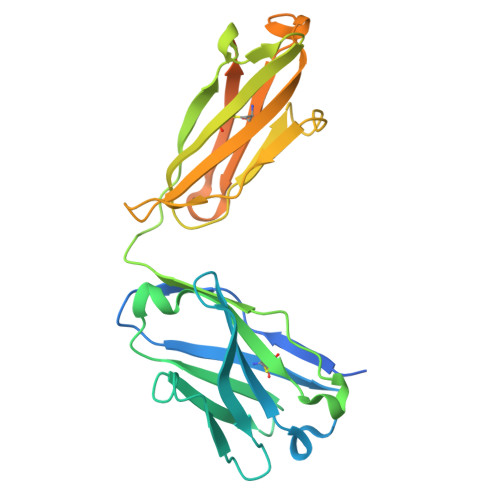
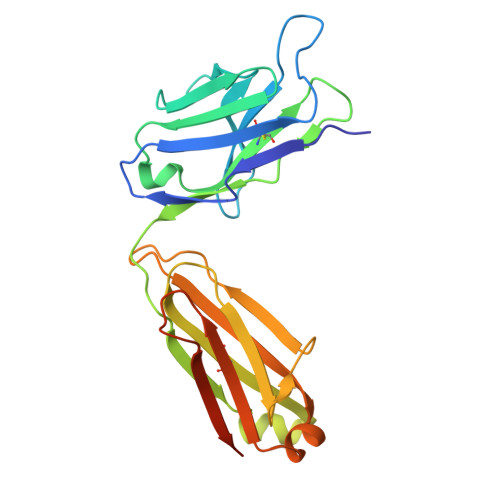
Switchbody, an Antigen-Responsive Enzyme Switch Based on Antibody and Its Working Principle.
Yasuda, T., Ueno, Y., Taguchi, M., Tochio, N., Yagi, H., Yazaki, S., Arai, R., Zhu, B., Kigawa, T., Ueda, H., Kitaguchi, T.(2025) Adv Sci (Weinh) 12: e08422-e08422
- PubMed: 40953287 Search on PubMed
- DOI: https://doi.org/10.1002/advs.202508422
- Primary Citation Related Structures:
9LUK - PubMed Abstract:
An enzyme switch, termed "Switchbody", is developed by fusing an antibody with a fragment of a split enzyme for the precise regulation of enzyme activity in response to an antigen. A luciferase-based Switchbody is engineered by fusing the NanoLuc luciferase fragment HiBiT to the N-terminus of an antibody. The enzyme activity of the Switchbody increases upon the addition of an antigen in a dose-dependent manner in the presence of the complementary fragment LgBiT and its substrate furimazine, demonstrating the potential of the luciferase-based Switchbody as a biosensor. As its working principle, ELISA shows that the interaction between HiBiT and LgBiT is facilitated by antigen binding. Moreover, X-ray crystallography and NMR reveal the heterogeneous trapped state of the HiBiT region and an increasing motility of HiBiT region upon antigen binding, respectively. MD simulations and luminescence measurements show that antigen disrupted the trapping of HiBiT in the antibody, enabling its release. By applying this "Trap and Release" principle to Protein M, an antibody-binding protein, label-free IgG antibodies are successfully converted into bioluminescent Switchbodies. This adaptable Switchbody platform has the potential to expand switching technology beyond luciferase to various other enzymes in the future.
- Laboratory for Chemistry and Life Science, Institute of Integrated Research, Institute of Science Tokyo, Yokohama, 226-8501, Japan.
Organizational Affiliation: