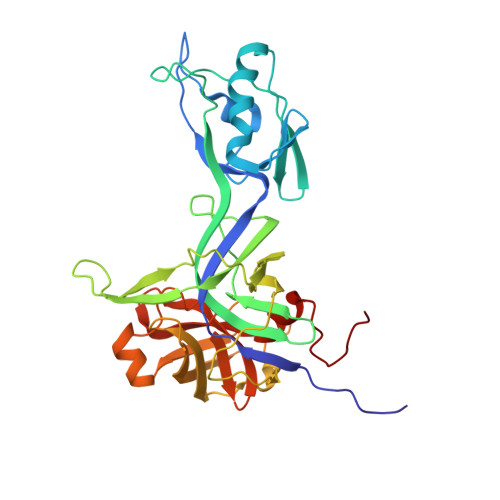
Structural characterization and biophysical analysis of recombinant SpoIVB variants: insights into PDZ and serine protease domain interactions.
Zhu, J., Zhang, X., Zhang, X., Sun, G., Xu, P., Yuan, C., Huang, M., Jiang, L.(2025) Microbiol Spectr 13: e0039825-e0039825
- PubMed: 40810511 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/spectrum.00398-25
- Primary Citation Related Structures:
9LNF - PubMed Abstract:
Sporulation factor IV B protease (SpoIVB), as a PDZ-protease, plays a central role in cellular differentiation via activating pro-σ K processing at the σ K checkpoint during spore formation. However, the molecular mechanism and structure of SpoIVB remain unclear. In this study, we expressed and characterized several recombinant variants of SpoIVB, including SpoIVB 75-426 , SpoIVB 75-426-S378A , and SpoIVB 101-426-S378A . Their structural properties were analyzed through dynamic light scattering, size-exclusion chromatography, small-angle X-ray scattering (SAXS), and X-ray crystallography. The crystal structure of SpoIVB 101-426-S378A was determined at 2.49 Å resolution, revealing a unique PDZ domain arrangement and an unusual catalytic triad in the serine protease domain. SAXS analysis demonstrated that SpoIVB 75-426-S378A adopts a monomeric form with a folded but flexible structure in solution, while the S378A mutation alters its hydrodynamic radius ( R H ) and overall compactness. These findings provide new insights into the structural dynamics of SpoIVB, including its monomeric state, PDZ domain interactions, and the functional implications of the S378A mutation. This study lays the groundwork for further investigations into the mechanistic role of SpoIVB in biological systems and its potential as a therapeutic target.IMPORTANCESporulation factor IV B protease (SpoIVB) is a pivotal PDZ-protease regulating the σ K checkpoint during bacterial sporulation, yet its structural and mechanistic details remain elusive. This study provides the first atomic-resolution crystal structure of a SpoIVB variant (SpoIVB 101-426-S378A ), revealing a non-canonical catalytic triad (His236-Ala378-Thr393) and a unique PDZ domain insertion into the serine protease core. Biophysical analyses demonstrate that the S378A mutation enhances structural compactness and monomeric stability, while small-angle X-ray scattering confirms flexibility in the N-terminal region. These findings challenge traditional views of serine protease mechanisms and unveil novel regulatory interactions between PDZ and catalytic domains. The structural insights advance understanding of SpoIVB's role in σ K activation and lay a foundation for targeting PDZ-protease interfaces in antibacterial strategies. This work bridges critical gaps in bacterial developmental biology and highlights SpoIVB as a potential therapeutic target.
- College of Chemistry, Fuzhou University, Fuzhou, Fujian, China.
Organizational Affiliation: