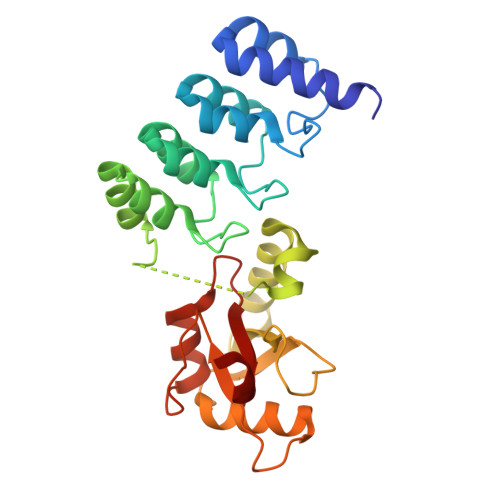
A non-canonical ARMS-GABARAP interaction modulates dendritic spine formation and synaptic development.
Jiang, W., Ye, J., Chen, J., Wang, X., Li, Y., Li, J., Mei, Y., Lyu, Y., Hu, W., Wang, C.(2026) EMBO J 45: 1109-1135
- PubMed: 41507394 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s44318-025-00669-w
- Primary Citation Related Structures:
9L9I - PubMed Abstract:
ARMS (ankyrin repeat-rich membrane spanning) is a scaffold protein essential for neurotrophic signaling, synaptic development, and cytoskeletal remodeling. Despite its central role in neuronal function, how ARMS is regulated at the molecular level remains poorly understood. Here, we identify GABARAP, an Atg8-family autophagy adaptor, as a novel ARMS-binding protein that directly interacts with its N-terminal ankyrin repeats. We present the crystal structure of the ARMS-GABARAP complex, revealing an atypical interaction mode distinct from canonical LIR-dependent Atg8 interactions. Remarkably, ARMS specifically binds to the GABARAP subfamily of Atg8 proteins, setting it apart from the LC3 subfamily. Functional analysis demonstrates that GABARAP negatively regulates ARMS-mediated dendritic spine development and maturation in hippocampal neurons. Additionally, disrupting the ARMS-GABARAP complex using ankyrin-derived peptides alters ARMS subcellular localization, increasing its accumulation in the soma of neurons. Collectively, our findings uncover a novel ARMS-GABARAP interaction mechanism, establish the regulatory role of this complex in neuronal protein homeostasis, and suggest potential therapeutic strategies for targeting scaffold protein interactions in neurodevelopmental and neurodegenerative disorders.
- Department of Neurology, the First Affiliated Hospital of USTC, Ministry of Education Key Laboratory for Membraneless Organelles and Cellular Dynamics, School of Life Sciences, University of Science and Technology of China, 230027, Hefei, China.
Organizational Affiliation: