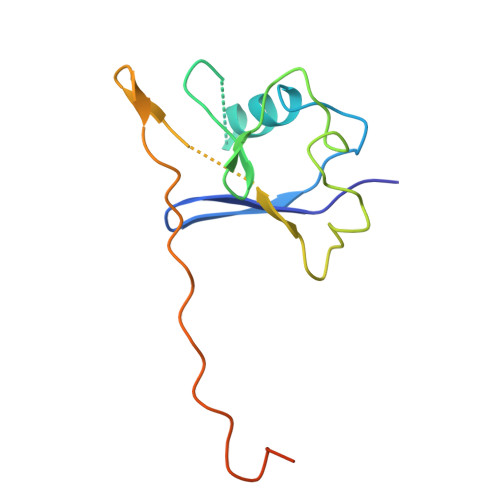
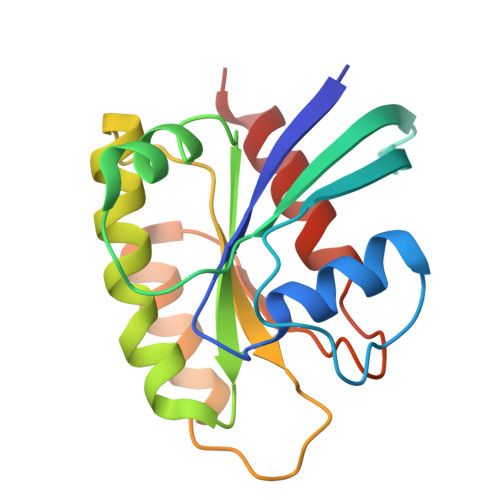
Discovery of KRAS(G12D) selective degrader ASP3082.
Yoshinari, T., Nagashima, T., Ishioka, H., Inamura, K., Nishizono, Y., Tasaki, M., Iguchi, K., Suzuki, A., Sato, C., Nakayama, A., Amano, Y., Tateishi, Y., Yamanaka, Y., Osaki, F., Yoshino, M., Kuramoto, K., Imaizumi, T., Hayakawa, M.(2025) Commun Chem 8: 254-254
- PubMed: 40849515 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42004-025-01662-4
- Primary Citation Related Structures:
9L6A, 9L6F - PubMed Abstract:
Kirsten rat sarcoma viral oncogene homolog (KRAS) is one of the most frequently mutated oncogenes in multiple cancers. Multiple types of KRAS mutation are observed in various patients with cancer, and the KRAS(G12D) mutation is the most common. Although multiple covalent inhibitors of the KRAS(G12C) mutation have been identified and clinically validated to date, no drugs have been approved yet for other mutations, including G12D. Herein, we report the discovery and characterization of ASP3082, a KRAS(G12D)-selective degrader, and the crystal structure of the drug-induced ternary complex of KRAS(G12D)/ASP3082/VHL (von Hippel-Lindau). We have also demonstrated an efficient structure-based rational optimization approach, which could be applicable for the optimization of other bifunctional proximity-inducing drugs. ASP3082 effectively induces KRAS(G12D) protein degradation with remarkable selectivity, demonstrates highly efficacious and durable pharmacological activity, and induces tumor regression in multiple KRAS(G12D)-mutated cancer xenograft models. Our results suggest that ASP3082 is a potential therapeutic agent for KRAS(G12D)-mutated cancer, and is now under clinical investigation.
- Engineered Small Molecules, Astellas Pharma Inc., Tsukuba, Japan. tomohiro.yoshinari@astellas.com.
Organizational Affiliation: