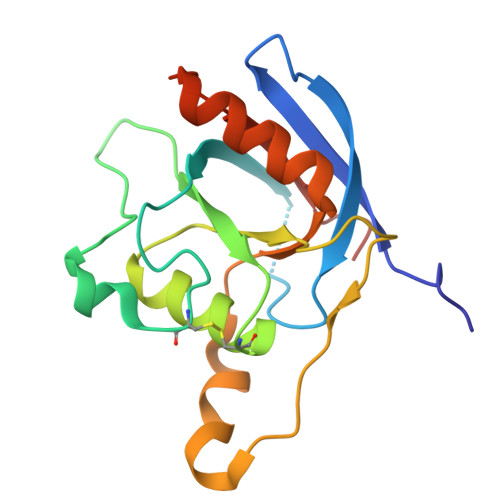
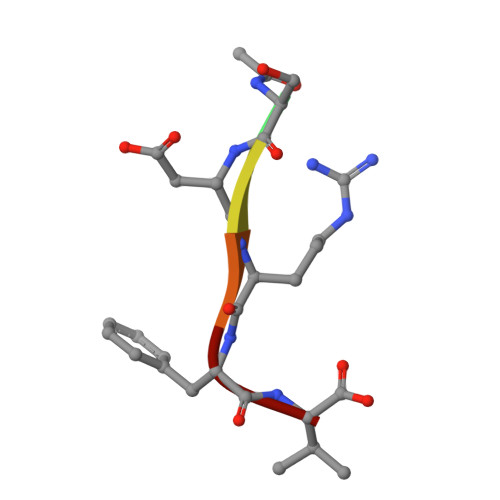
Structural insights into how vacuolar sorting receptor recognizes the C-terminal sorting determinant of a vicilin-like seed storage protein.
Lui, S.N., Tsao, H.E., Lo, A.H., Jiang, L., Wong, K.B.(2026) FEBS J 293: 257-270
- PubMed: 40891502 Search on PubMed
- DOI: https://doi.org/10.1111/febs.70245
- Primary Citation Related Structures:
9L4O, 9L4P - PubMed Abstract:
During seed development, vacuolar sorting receptors (VSRs) recognize a sequence-specific vacuolar sorting determinant located at the C terminus (ctVSD) of storage proteins, thereby sorting them into protein storage vacuoles. The protease-associated (PA) domain of VSRs is responsible for interacting with the ctVSD of cargo proteins. Here, we report the crystal structure of the PA domain of Arabidopsis vacuolar-sorting receptor 1 (VSR1) in complex with the C-terminal pentapeptide ( 507 SDRFV 511 ) of vicilin-like seed storage protein 22 (VL22). Structural comparison with the apo form of VSR1 reveals conformational changes in four switch regions in the PA domain. VL22 binds to a cradle of VSR1 formed by residues in the cargo-binding loop, the switch I and III regions. The C-terminal carboxyl group of VL22 is recognized by forming salt bridges with the invariant Arg95 of VSR1. Compared with the structure of VSR1-PA in complex with the ctVSD of cruciferin 1, VL22 makes extra hydrophobic interactions with the cargo-binding loop and hydrogen bonds with switch I residues in VSR1. Tagging the C-terminal sequence of VL22, but not VL22-R509P, VL22-V511P, VL22-R509P-V511P nor vicilin-like seed storage protein 43 (VL43), redirected secretory red fluorescent protein (spRFP) to the vacuoles in Arabidopsis protoplasts. Scanning mutagenesis identified an E519S substitution converting the C-terminal sequence of VL43 to a sorting determinant that can redirect spRFP to the vacuoles, suggesting that charge-charge repulsion prevents the receptor-cargo interactions between VL43 and VSR1. The recognition of ctVSD by VSRs is likely promiscuous, resulting from the additive effect of individual preference of residues in the ctVSD.
- Centre for Protein Science and Crystallography, State Key Laboratory of Agrobiotechnology, School of Life Sciences, The Chinese University of Hong Kong, Hong Kong, China.
Organizational Affiliation: