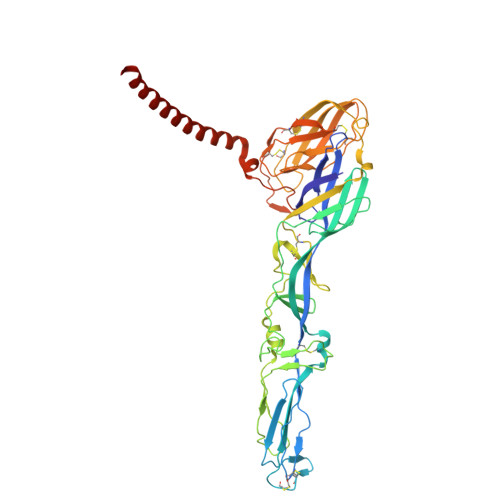
Structural basis for engagement of Western Equine Encephalitis Virus with the PCDH10 receptor.
Liang, S., Yang, Y., Liu, Y., Xu, Z., Hou, J., Li, D., Zhao, L., Hu, C., Liu, X., Rao, Z., Wang, Y., Lou, Z.(2025) Nat Commun 16: 6290-6290
- PubMed: 40628733 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-61659-4
- Primary Citation Related Structures:
9L3V, 9L41 - PubMed Abstract:
PCDH10 is a newly identified general receptor for Western equine encephalitis virus (WEEV) members, a group of encephalitic alphaviruses that cause severe diseases in humans and equids. While WEEV typically binds PCDH10 as a receptor, nonpathogenic strains have evolved to lose mammalian PCDH10 binding, retaining only avian PCDH10 affinity. Virulent strains also engage VLDLR and ApoER2 as alternative receptors. Here, we determine the structure of WEEV strain 71V1658 virus-like particles (VLPs) in complex with human PCDH10 extracellular cadherin repeats 1-2 (EC1-EC2) by cryo-electron microscopy at 2.99 Å resolution. EC1 inserts into a cleft clamped by two adjacent E2-E1 heterodimers within a single trimeric spike, whereas EC2 maintains no contact with the WEEV VLP. Mutagenesis studies elucidate the impacts of the interacting residues on PCDH10. And residue 153 of E2 is crucial for PCDH10 binding, and the E2 Q153L mutation observes in the nonpathogenic strain Imperial-181 restores its ability to bind to PCDH10. Moreover, the arginine residue at position 89 on avian PCDH10 is essential for its interaction with strain Imperial-181. These results advance our understanding of receptor recognition by alphaviruses and the shift in receptor usage, providing insights for the development of antiviral therapies.
- School of Basic Medical Sciences, Tsinghua University, Beijing, China.
Organizational Affiliation: