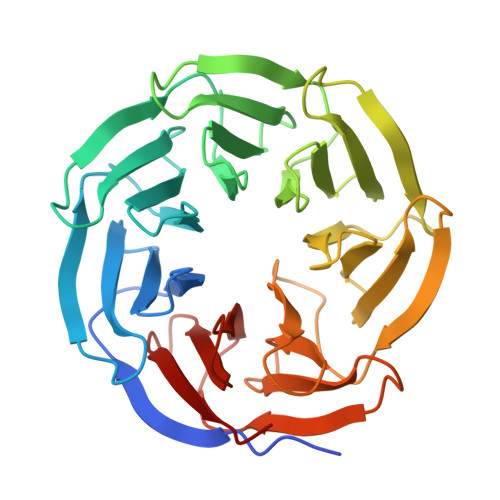
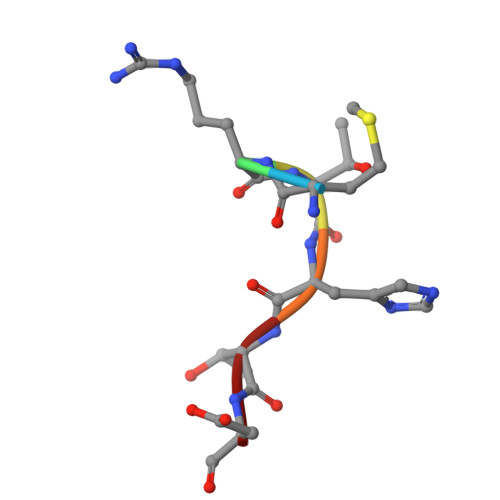
Structural characterization of endogenous microprotein EMBOW reveals an alternative MRT motif for WDR5-interacting site recognition.
Yang, Y., Pan, Y., Zhang, S., Wang, H., Sun, X., Hang, T., Xu, L.(2026) FEBS J
- PubMed: 41689289
- DOI: https://doi.org/10.1111/febs.70450
- Primary Citation of Related Structures:
9KFM, 9KFN, 9KFO, 9L0T, 9L0V - PubMed Abstract:
WD repeat-containing protein 5 (WDR5) is a conserved chromatin regulator that engages numerous binding partners via a central arginine-binding pocket known as the WDR5-interacting (WIN) site. Endogenous microprotein binder of WDR5 (EMBOW, also known as SCRIB overlapping open reading frame protein), recently identified as an endogenous WDR5 interactor, lacks the canonical [ACR]-R-[TASCK] WIN motif, and its mode of recognition remains unknown. Here, we present the 1.80 Å crystal structure of WDR5 in complex with an EMBOW-derived peptide. Our structural analysis reveals that EMBOW engages the WIN site through a Met1-Arg2-Thr3 (MRT) triad. The bulky Met1 residue occupies the conserved WIN site pocket, and mutation of Thr3 to valine reduces binding affinity, while N-terminal Gly-Ser insertion preserves binding, indicating a degree of structural tolerance. Binding assays and mutational analysis underscore the functional importance of the MRT triad. Furthermore, structural and biochemical studies of MRT-containing peptides from RNA-binding protein 15 (RBM15) and zinc finger and SCAN domain-containing protein 10 (ZSCAN10) suggest that this motif may serve as an alternative WIN site recognition signature. In summary, our findings define the molecular basis of EMBOW-WDR5 interaction and expand the sequence space compatible with WIN site engagement.
- School of Life Sciences, Anhui University, Hefei, China.
Organizational Affiliation: