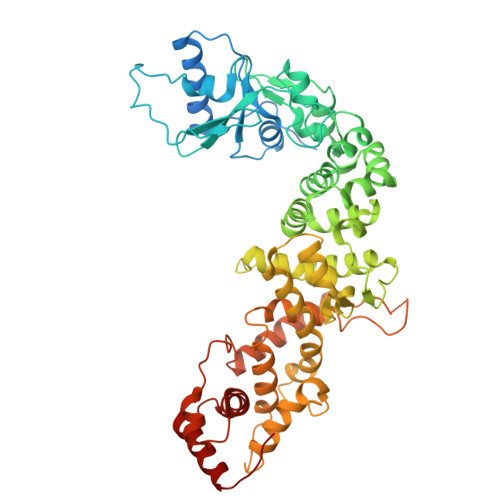
Crystal structure of the CCA-adding enzyme from Arabidopsis thaliana .
Wang, X., Li, Y.Y., Dou, Z.Y., Wang, J., Liu, L.(2025) J Struct Biol X 11: 100127-100127
- PubMed: 40519583 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.yjsbx.2025.100127
- Primary Citation Related Structures:
9KMA - PubMed Abstract:
The 3'-terminal CCA-end of tRNA is essential for the attachment of amino acids and correct positioning of the aminoacyl-tRNA in the ribosome. In higher plants, the CCA sequence is synthesized, maintained, and repaired by class-II CCA-adding enzymes encoded by a single nuclear gene but multi-targeted to the nucleus, cytoplasm, plastids, and mitochondria. The structure of plant class-II CCA-adding enzyme remains unsolved. Here we describe the crystal structure of CCA-adding enzyme from Arabidopsis thaliana ( At CCA) . The overall structure of At CCA is similar to other class-II CCA-adding enzymes , but significant differences occur in the body domain. Structural comparison of body and tail domains between At CCA and other class-II CCA-adding enzymes unravels three specific regions of At CCA. Based on the modeled At CCA-tRNA complex, At CCA may have a different tRNA binding pattern. The three specific regions located in the body domain of At CCA also provide candidate regions for multi-targeted sorting.
- School of Life Sciences, Anhui University, Hefei, Anhui 230601, China.
Organizational Affiliation: