Structure of KATP in complex with SpTx1
Chen, L., Wang, M.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| ATP-sensitive inward rectifier potassium channel 11 | 390 | Homo sapiens | Mutation(s): 2 Gene Names: KCNJ11 | 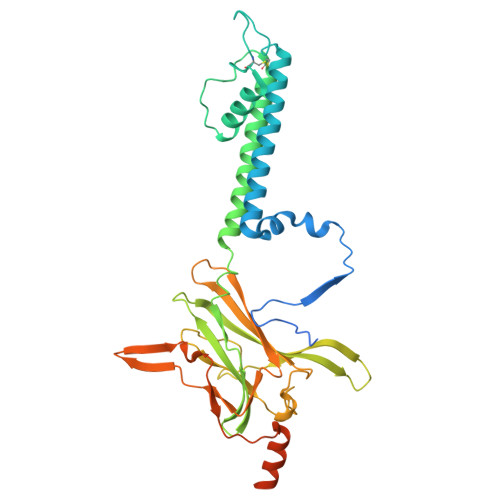 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for Q14654 (Homo sapiens) Explore Q14654 Go to UniProtKB: Q14654 | |||||
PHAROS: Q14654 GTEx: ENSG00000187486 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q14654 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| ATP-binding cassette sub-family C member 8 isoform X2 | 1,582 | Mesocricetus auratus | Mutation(s): 0 Gene Names: Abcc8 |  | |
UniProt | |||||
Find proteins for A0A1U7R319 (Mesocricetus auratus) Explore A0A1U7R319 Go to UniProtKB: A0A1U7R319 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | A0A1U7R319 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| SpTx1 | 56 | Scolopendra polymorpha | Mutation(s): 0 | 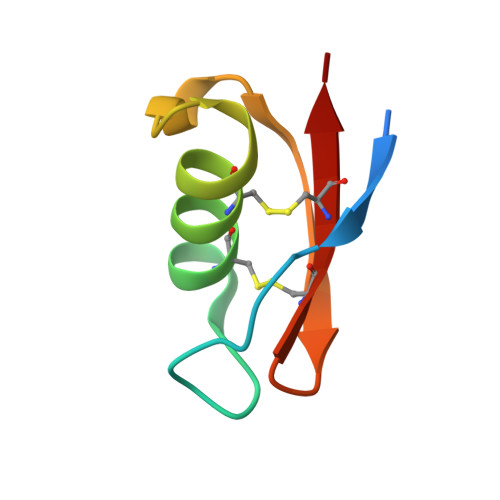 | |
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| ATP Query on ATP | J [auth A] L [auth B] M [auth C] O [auth D] P [auth E] | ADENOSINE-5'-TRIPHOSPHATE C10 H16 N5 O13 P3 ZKHQWZAMYRWXGA-KQYNXXCUSA-N |  | ||
| GBM Query on GBM | K [auth B], N [auth D], Q [auth F], T [auth H] | 5-chloro-N-(2-{4-[(cyclohexylcarbamoyl)sulfamoyl]phenyl}ethyl)-2-methoxybenzamide C23 H28 Cl N3 O5 S ZNNLBTZKUZBEKO-UHFFFAOYSA-N |  | ||
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Natural Science Foundation of China (NSFC) | China | 91957201 |