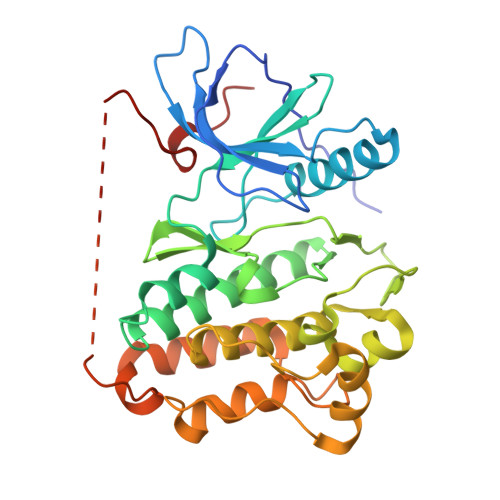
Crystal structure of EGFR T790M/C797S/L858R mutant in complex with 2,2-dichloro-N-(5-((5-chloro-4-((2-(dimethylphosphoryl)phenyl)amino)pyrimidin-2-yl)amino)-4-methoxy-2-(4-(4-methylpiperazin-1-yl)piperidin-1-yl)phenyl)acetamide
Wang, Y., Ma, L., Yang, X.To be published.