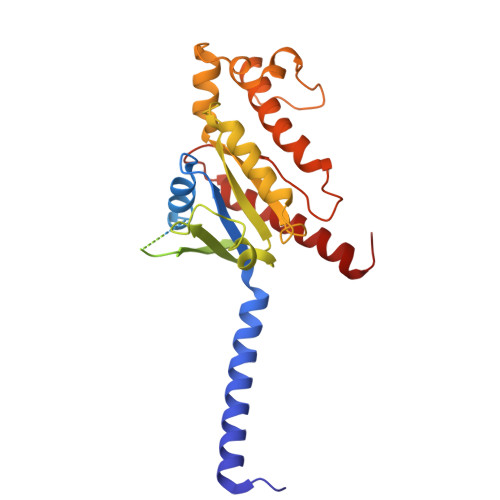
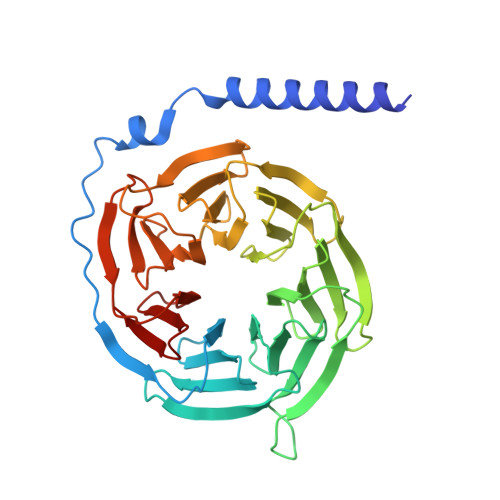
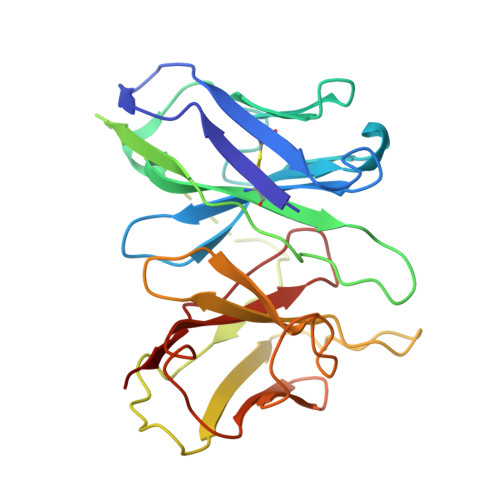
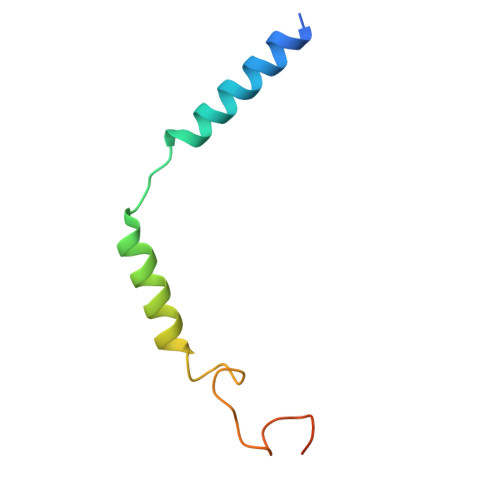
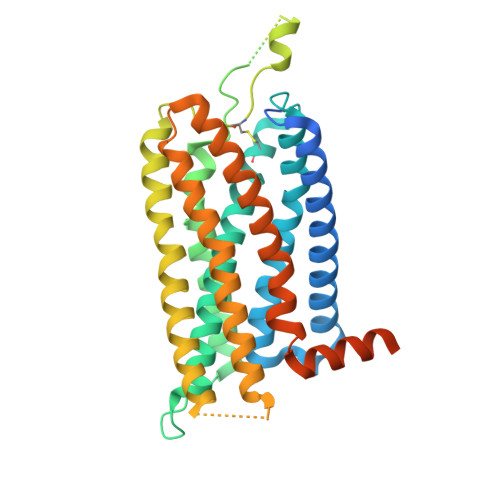
Decoding the structural basis of ligand recognition and biased signaling in the motilin receptor.
You, C., Jiang, M., Gao, T., Zhu, Z., He, X., Xu, Y., Gao, Y., Jiang, Y., Xu, H.E.(2025) Cell Rep 44: 115329-115329
- PubMed: 39987561 Search on PubMed
- DOI: https://doi.org/10.1016/j.celrep.2025.115329
- Primary Citation Related Structures:
9JMC, 9JMD - PubMed Abstract:
The motilin receptor (MTLR) is a key target for treating gastrointestinal (GI) disorders like gastroparesis, yet developing effective agonists remains challenging due to drug tolerance and signaling bias. We present cryoelectron microscopy (cryo-EM) structures of MTLR bound to azithromycin, a macrolide antibiotic, and DS-3801b, a non-macrolide agonist. Distinct ligand recognition mechanisms are revealed, with azithromycin binding deeply within the orthosteric pocket and DS-3801b adopting a special clamp-like conformation stabilized by a water molecule. We also highlight the critical role of extracellular loop 2 (ECL2) in ligand specificity and signaling pathway activation, affecting both G-protein and β-arrestin signaling. Additionally, the "D 2.60 R 2.63 S 3.28 " motif and interactions around transmembranes 6/7 (TM6/7) are identified as key drivers of signaling selectivity. These findings offer insights into the structural dynamics of MTLR, laying the groundwork for the rational design of next-generation GI prokinetic drugs with enhanced efficacy and safety.
- The State Key Laboratory of Drug Research, Shanghai Institute of Materia Medica, Chinese Academy of Sciences, Shanghai 201203, China; University of Chinese Academy of Sciences, Beijing 100049, China. Electronic address: youchongzhao@gmail.com.
Organizational Affiliation: