An Engineered Soluble Single-Chain TCR Engager for KRAS-G12V Specific Tumor Immunotherapy.
Ma, K., Wang, J., Jiang, M., He, J., Li, F., Lu, D., Su, C., Chai, Y., Jin, W., Chen, Y., Zhang, C.W.H., Ma, X., Tan, H., Gao, G.F., Tan, S.(2025) Adv Sci (Weinh) 12: e00181-e00181
- PubMed: 40470869 Search on PubMed
- DOI: https://doi.org/10.1002/advs.202500181
- Primary Citation Related Structures:
9IKY - PubMed Abstract:
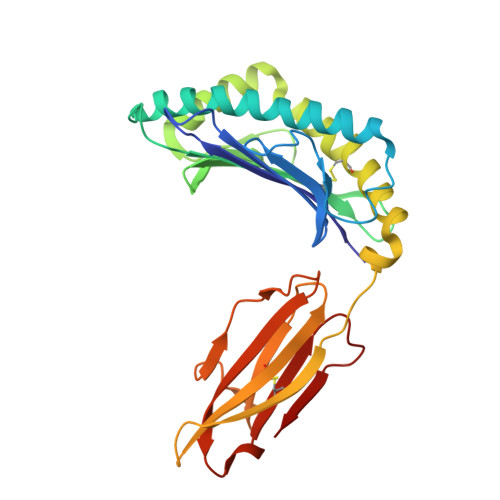
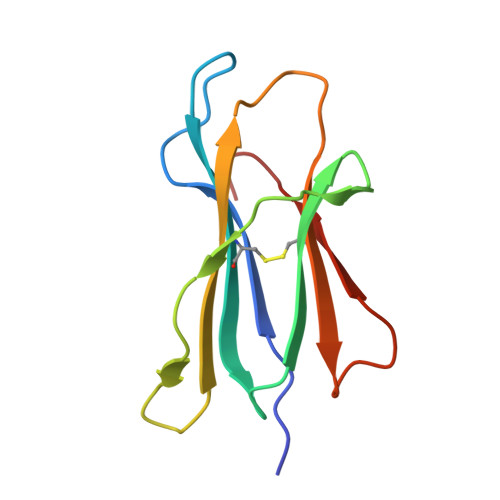
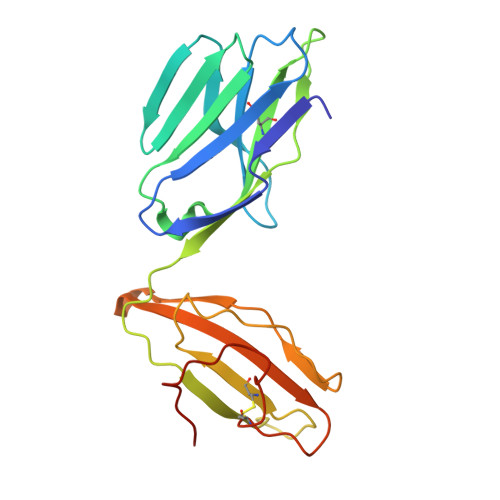
T cell receptor (TCR) based immunotherapy is an attractive strategy to target a wide range of intra-tumoral antigens and elicit robust tumor cytotoxicity. However, engineering soluble TCR engagers that preserve physiological affinity is crucial for universal TCR drug development, yet remains challenging. In the present study, multiple TCR engagers featuring diverse architectures based on the KRAS-G12V specific 1-2C TCR in the context of HLA-A*11:01 is designed and evaluated. Notably, a soluble tandem double single-chain TCR (STanD-scTCR) engager, comprising two repeated single-chain variable fragment (scFv) TCRs, exhibit enhanced binding avidity and potent T-cell activation. Through site-directed mutagenesis, T96F mutation (T96F-TCR) within the TCR β chain is identified, which substantially augment T cell reactivity while maintaining physiological affinity and minimizing off-target cross-reactivity. The T96F-mutated STanD-scTCR engager demonstrates improved antigen sensitivity, promotes multi-functional T-cell responses, and facilitates immune synapse formation between T cells and target cells. In a xenograft tumor model harboring the KRAS-G12V mutation, the TCR engager displays substantial tumor suppression efficacy. These findings underscore the therapeutic potential of 1-2C STanD-scTCR engage in targeting KRAS-G12V mutations in the context of HLA-A*11:01. Furthermore, the engineering strategies employ in the development of STanD-scTCR engager provide an invaluable for future designs of TCR engager drugs.
- Department of Infectious Diseases, Shenzhen Children's Hospital, Shenzhen, 518026, China.
Organizational Affiliation: