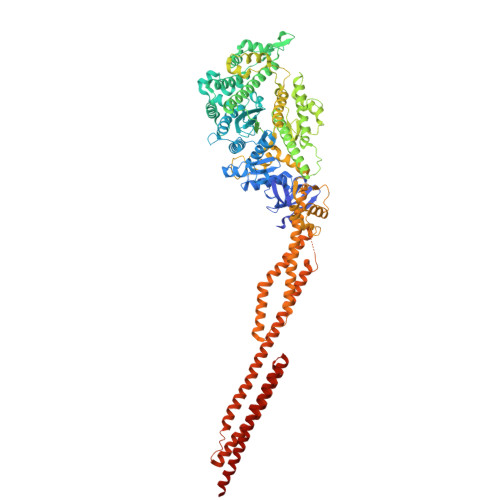
Structure of the human nonmuscle myosin 2A motor domain: Insights into isoform-specific mechanochemistry.
Heiringhoff, R.S., Greve, J.N., Zahn, M., Manstein, D.J.(2025) J Biological Chem 301: 110691-110691
- PubMed: 40939649 Search on PubMed
- DOI: https://doi.org/10.1016/j.jbc.2025.110691
- Primary Citation Related Structures:
9IHL - PubMed Abstract:
Non-muscle myosin 2A (NM2A) is the predominant myosin isoform in non-muscle cells. Together with its paralogues NM2B and NM2C, NM2A enables tension and force generation, driving essential cellular processes such as membrane protrusion and retraction, directed migration, adhesion and cytokinesis. The NM2 isoforms display paralogue-specific mechanochemical characteristics that support their specific cellular functions. Here, we determined the structure of the human NM2A motor domain, addressing a critical gap in understanding myosin family diversification. Based on our experimentally resolved 2.1 Å structure of the NM2A motor domain in its nucleotide-free state, we demonstrate, through integrative modeling of NM2-actin complexes and molecular dynamics simulations, how sequence differences between NM2A and NM2B underpin their functional specialization. Loop2 emerges as a critical determinant of isoform-specific behavior. Comparative analysis of ATP interaction fingerprints across NM2 isoforms reveals a conserved ATP binding mechanism. These findings illuminate an allosteric energy transduction pathway that connects sequence variation to actin-binding dynamics, providing mechanistic insight into isoform-specific cytoskeletal functions.
- Institute for Biophysical Chemistry, Fritz-Hartmann-Centre for Medical Research, Hannover Medical School, 30625 Hannover, Germany,; Division for Structural Biochemistry, Hannover Medical School, 30625 Hannover, Germany.
Organizational Affiliation: