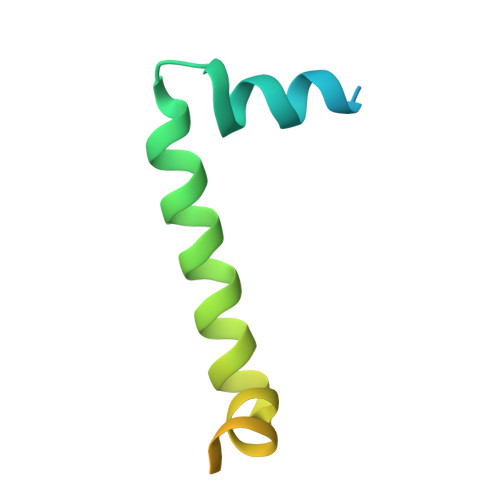
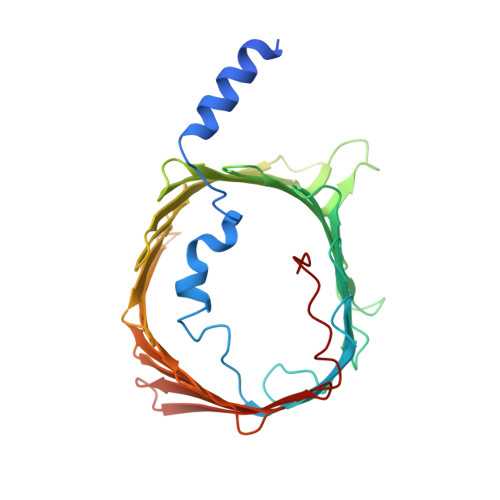
Structures of Chaetomium thermophilum TOM complexes with bound preproteins.
Agip, A.A., Ornelas, P., Yang, T.J., Uboldi, E., Hader, S., McDowell, M.A., Kuhlbrandt, W.(2025) Proc Natl Acad Sci U S A 122: e2507279122-e2507279122
- PubMed: 40674418 Search on PubMed
- DOI: https://doi.org/10.1073/pnas.2507279122
- Primary Citation Related Structures:
9I6B, 9I7P, 9I7S, 9I7T - PubMed Abstract:
Mitochondria import most of their proteins from the cytoplasm through the TOM complex. Preproteins containing targeting signals are recognized by the TOM receptor subunits and translocated by Tom40 across the outer mitochondrial membrane. We present four structures of the preprotein-bound and preprotein-free TOM core and holo complexes from the thermophilic fungus Chaetomium thermophilum , obtained by single-particle electron cryomicroscopy. Our structures reveal the symmetric arrangement of two copies of the Tom20 receptor subunit in the TOM holo complex. Several different conformations of Tom20 within the TOM holo complex highlight the dynamic nature of the receptor. The structure of preprotein-bound Tom20 provides insight into the early stages of protein translocation.
- Department of Structural Biology, Max Planck Institute of Biophysics, Frankfurt 60438, Germany.
Organizational Affiliation: