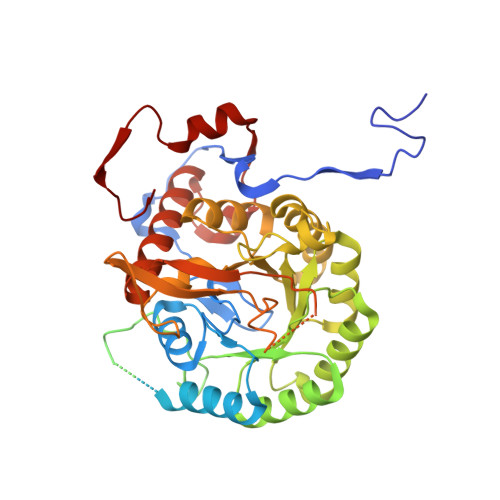
Conformational landscape of the mycobacterial inosine 5'-monophosphate dehydrogenase octamerization interface.
Bulvas, O., Knejzlik, Z., Filimonenko, A., Kouba, T., Pichova, I.(2025) J Struct Biol 217: 108198-108198
- PubMed: 40107326 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2025.108198
- Primary Citation Related Structures:
9I0K, 9I0L, 9I0M - PubMed Abstract:
Inosine 5'-monophosphate dehydrogenase (IMPDH), a key enzyme in bacterial purine metabolism, plays an essential role in the biosynthesis of guanine nucleotides and shows promise as a target for antimicrobial drug development. Despite its significance, the conformational dynamics and substrate-induced structural changes in bacterial IMPDH remain poorly understood, particularly with respect to its octameric assembly. Using cryo-EM, we present full-length structures of IMPDH from Mycobacterium smegmatis (MsmIMPDH) captured in a reaction intermediate state, revealing conformational changes upon substrate binding. The structures feature resolved flexible loops that coordinate the binding of the substrate, the cofactor, and the K + ion. Our structural analysis identifies a novel octamerization interface unique to MsmIMPDH. Additionally, a previously unobserved barrel-like density suggests potential self-interactions within the C-terminal regions, hinting at a regulatory mechanism tied to assembly and function of the enzyme. These data provide insights into substrate-induced conformational dynamics and novel interaction interfaces in MsmIMPDH, potentially informing the development of IMPDH-targeted drugs.
- Institute of Organic Chemistry and Biochemistry of the Czech Academy of Sciences, Prague, Czech Republic.
Organizational Affiliation: