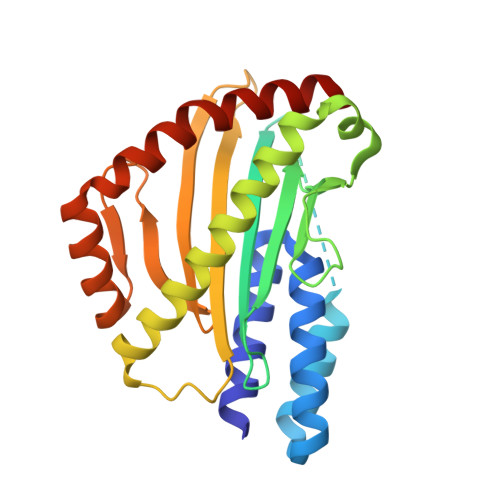
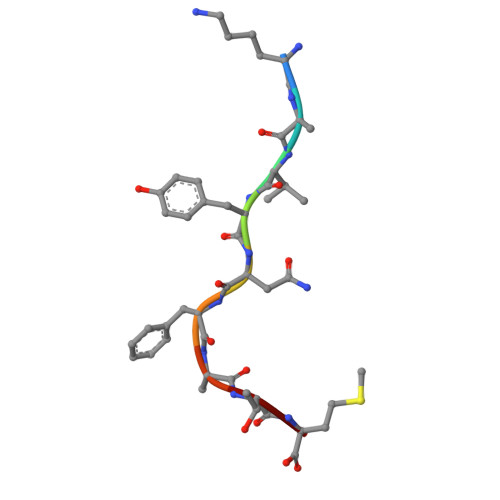
Design of solubly expressed miniaturized SMART MHCs.
White, W.L., Bai, H., Kim, C.J., Jude, K.M., Sun, R., Guerrero, L., Han, X., Chen, X., Chaudhuri, A., Bonzanini, J.E., Sun, Y., Onwuka, A.E., Wang, N., Wang, C., Nygren, P.A., Li, X., Goreshnik, I., Allen, A., Levine, P.M., Kueh, H.Y., Jewett, M.C., Sgourakis, N.G., Achour, A., Garcia, K.C., Baker, D.(2026) Proc Natl Acad Sci U S A 123: e2505932123-e2505932123
- PubMed: 41481462 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2505932123
- Primary Citation Related Structures:
9HY4, 9NDS - PubMed Abstract:
The precise recognition of specific peptide-major histocompatibility complex (pMHC) complexes by T cell receptors (TCRs) plays a key role in infectious disease, cancer, and autoimmunity. A critical step in many immunobiological studies is the identification of T cells expressing TCRs specific to a given pMHC antigen. However, the intrinsic instability of empty class-I MHCs limits their soluble expression in Escherichia coli and makes it very difficult to characterize even a small fraction of possible pMHC/TCR interactions. To overcome this limitation, we designed small proteins which buttress the peptide binding groove of class I MHCs, replacing β2-microglobulin (β2m) and the heavy chain α3 domain, and enable soluble and partially soluble expression in E. coli of H-2D b and A*02:01, respectively. We demonstrate that these soluble, monomeric, antigen-receptive, truncated (SMART) MHCs retain both peptide- and TCR-binding specificity and that peptide-bound structures of both allomorphs are similar to their full-length, native counterparts. With extension to the majority of HLA alleles, SMART MHCs should be broadly useful for probing the T cell repertoire in approaches ranging from yeast display to T cell staining.
- Department of Bioengineering, University of Washington, Seattle, WA 98195.
Organizational Affiliation: